大约5亿年前绿色植物登陆, 从根本上改变了地表碳、水和营养元素的生物地球化学循环(Brodribb & McAdam, 2017)。尽管陆生和水生植物的栖息环境差别很大, 植物出现了气孔, 其光合作用、水分运输、CO2固定/O2消耗等功能演化更复杂高效, 但在过去的研究中观察到进化层级较高的种子植物与水生植物藻类均有光合作用昼夜节律和光周期反应。昼夜节律对多种植物生物学功能具有重要作用, 如气孔开放、光合作用、叶片运动、新陈代谢、开花诱导、植物生长和胁迫响应(Dodd et al., 2014; Greenham & McClung, 2015)。过去50年内的大量文献记录了植物气体交换的昼夜节律调节作用, 但这些文献集中在少数模式物种, 并主要描述昼夜光照改变的环境条件下的指标变化。此前的研究表明温度和叶-气水汽压差(VPD)对气体交换调节起到主要作用; 基于C3和C4植物的meta分析发现, 昼夜节律可以解释净光合速率(Pn)和气孔导度(Gs)日变化的15%-25%和30%-35% (Resco de Dios & Gessler, 2018), 昼夜节律对气体交换的调节作用甚至可以匹敌上述两种环境因素; 昼夜节律的调节作用已经用于植被生产力和全球变化模型(Resco de Dios et al., 2013, 2016), 具有重要的生态意义。
生物钟是一种内源性的计时机制, 它根据接收的环境信号来预测未来的环境变化, 并在基因表达和生化调节上做好准备, 以此调节植物生理和新陈代谢的节律, 从而控制植物对环境信号的响应。基于拟南芥(Arabidopsis thaliana)的研究发现植物生物钟系统最重要的部分是核心振荡器。核心振荡器是一个转录因子调控网络, 由3个反馈环组成。第一个是核心反馈环, 由2个Myb类转录因子CCA1 (CIRCADIAN-CLOCK-ASSOCIATED 1)/LHY (LATE ELONGATED HYPOCOTYL)和PRR (PSEUDO RESPONSE REGULATOR)家族转录因子TOC1 (TIMING OF CAB 1)构成, CCA1/LHY与TOC1相互抑制。第二个循环是早间循环, CCA1/LHY一起正调节PRR7和PRR9的表达, 同时负调节PRR5基因, 而PRRs也可以负反馈调节CCA1和LHY。第三个循环是晚间循环, 由REVEILLE (RVEs)基因激活, 由LUX转录因子、ELF3/ELF4 (EARLY FLOWERING 3/4)蛋白形成EC复合体, 它抑制PRRs基因家族。此外, TOC1的稳定性受E3泛素连接酶GIGANTEA (GI)和ZEITLUPE (ZTL)的调节, 进而调节CCA1/LHY的表达。
尽管在拟南芥、水稻(Oryza sativa)等陆生高等植物和藻类等水生植物中对植物的节律调节作用有了较深入的研究; 然而, 从水生到陆生植物的演化类群中, 大部分植物类群的节律作用尚未研究。藓类作为第一批登陆的陆生植物, 其光合作用、水分利用等植物功能介于陆生和水生植物之间, 在陆地植物节律调节的进化研究方面具有重要的研究价值; 尤其是水分和温度等栖息环境发生重要变化, 藓类能否保持藓类植物光合固碳的节律作用? 另一方面, 过去的研究认为藓类作为缺乏维管束的非等水植物, 体内生理活动对周围环境的水分和光照响应更为灵敏, 难以像种子植物保持稳态; 因此本研究预测藓类植物的昼夜节律响应弱于种子植物。在藓类的节律响应研究方面, 前人对小立碗藓(Physcomitrella patens)16 h/8 h昼夜光周期处理下的转录组进行分析, 然而其结果大部分可以归属于对光照的响应而不是内在的节律响应。因此, 本研究对3种藓类进行15天12 h光照/12 h黑暗光照处理, 使其建立12 h/12 h昼夜调节的内在节律, 随之进行24 h/0 h全光照处理, 分析其在上述条件下气体交换内在节律的延续调节, 从而排除光照的影响; 进而对其中一种藓类进行NSC含量和转录组分析, 厘清光合作用和气孔调节相关基因表达的节律响应特点。
1 材料和方法
1.1 研究材料
本研究采用3种藓类植物, 分别为近危种脆枝青藓(Brachythecium thraustum)和广布种大灰藓(Hypnum plumaeforme)、长叶提灯藓(Mnium lycopodioides)。后两种藓类分布在中国大部分气候带(中温带、暖温带及亚热带), 因此可降低地区差异影响; 前者具有濒危特性, 对气候条件比较敏感; 因此这3种植物可以概括藓类植物的节律特点。藓类植物在西南科技大学植物荫棚中培养, 日平均气温为19-30 ℃, 日平均相对湿度为40%-60%。
1.2 研究方法
3种藓类于夏季移至人工气候室内驯化15天, 使其适应环境条件并重置周期和相位(生物钟的状态与24 h周期的位置关系)。人工气候室内光周期为12 h光照/12 h黑暗。光照时人工气候室光照强度500 μmol·m−2·s−1, 温度30 ℃, 相对湿度57%-60%、CO2浓度400 μmol·mol-1; 黑暗时人工气候室光照强度0 μmol·m−2·s−1, 温度19 ℃, 相对湿度80%, CO2浓度400 μmol·mol-1。驯化完成后转移到光照和温度恒定的人工气候室内(光照和温度与驯化期间昼间一致, 500 μmol·m−2·s−1, 30 ℃)进行 24 h/0 h全光照处理2天。为研究体内水分耗散和光合作用的节律变化, 对3种藓类驯化第15天(驯化最后一天)和第16、17天(两天全光照处理)的光合气体交换参数进行测定, 间隔4 h测量一次; 并测量全光照处理最后一天的NSC含量, 间隔时间同前。为进一步研究固碳耗水过程相关基因的节律变化, 对具有光合作用、蒸腾速率节律较强的脆枝青藓进行时序转录组表达及相关性分析。
1.3 测定指标与方法
1.3.1 光合气体交换参数及NSC含量
采用LI-6800光合测量系统(LI-COR, Lincoln, USA)监测3种藓类驯化第15天(驯化最后一天)和第16、17天(两天全光照处理)全光照处理的气体交换参数, 包括: Pn、蒸腾速率(Tr)。测定时间从8:00开始, 空气流速500 µmol·s-1, 叶室温度、湿度和光照强度设置与人工气候室条件一致; 每隔4 h测量一次, 叶组织采集后立即在液氮中冷冻然后放置在超低温冰箱保存备用。另采集叶组织, 使用微波炉快速杀青后于60 ℃烘箱干燥48 h至恒质量, 用于测定可溶性糖和淀粉含量(蒽酮比色法测定)。
1.3.2 转录组表达与相关性分析
为进一步研究固碳耗水过程相关基因的节律变化, 对具有光合作用、蒸腾速率节律较强的脆枝青藓进行转录组表达分析。在全光照处理的第二天(LL2d)每隔4 h对24 h/0 h处理的脆枝青藓采一次样品(采样时间同气体交换参数测定时间), 取6个时间段共18个样品。将脆枝青藓叶组织研磨成细粉, 提取RNA后构建cDNA文库, 获得18个cDNA文库(每个时间段含3次生物学重复); 进行Illumina测序。经过原始数据过滤、测序错误率检查、GC含量分布检查, 共获得51.5 Gb后续分析使用的Clean Data, 各样品Clean Data均达到6.12 Gb, Q30碱基百分比在94.65%以上, 测序质量和样品重复性良好, 满足分析要求。
原始数据过滤后得到高质量的clean reads, 使用Trinity (Grabherr et al., 2011)组装转录本, 采用RSEM (Li & Dewey, 2011)软件得到unigene的相对表达量(expected number of Fragments Per Kilobase of transcript sequence per Millions base pairs sequenced, FPKM)。脆枝青藓基因注释由Mercator4注释, 使用MapMan作图(Schwacke et al., 2019), 由R v4.1.1软件进行数据分析。直系同源基因采用OrthoFinder v2.5.4 (Emms & Kelly, 2019)和Diamond (Buchfink et al., 2015)分析获得; 参数如下: blastp中使用标准E < 10-10, query coverage > 50%和pident > 50%, 以避免在orthofinder进行比较分析时出现潜在偏差。采用JTK软件判断昼夜节律显著的基因。
1.4 数据分析
气体交换指标和NSC相关数据使用R软件进行处理、统计分析和绘图。对Pn、Tr使用广义相加模型(GAM)拟合, 基因表达热图使用complete-linkage聚类法进行聚类, 用R包“factoextra”对气孔运动相关节律性基因、Pn、 Tr进行主成分分析, 然后采用MapMan 4软件将相关性分析结果导入光合和碳固定过程进行绘图。
2 结果
2.1 藓类的Pn、Tr及NSC含量受昼夜节律调控
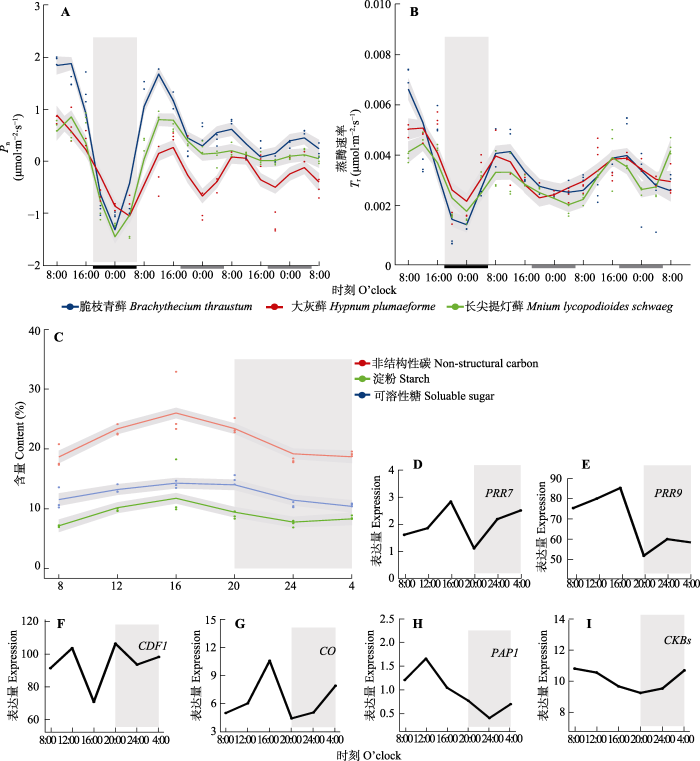
3种藓类经过正常光周期(12 h/12 h)适应15天后的Pn (夜间无光照情况下表现为呼吸速率)均出现昼夜节律振荡(图1A), 但是种间差异明显, 其中脆枝青藓振幅最强而长叶提灯藓最弱。通过GAM拟合估算正常光周期条件下脆枝青藓的Pn从夜间最低-1.29 µmol·m−2·s−1到白天最高1.89 µmol·m−2·s−1。在恒定光照条件下(LL, 24 h/0 h)第1天(LL1d)脆枝青藓的Pn振荡范围为0.32-1.69 µmol·m−2·s−1, 是正常光周期的43.1%, 第2天(LL2d)脆枝青藓的Pn振幅占正常光周期的16.7%。同样, 大灰藓LL1d的振幅占正常光周期的48.2%, LL2d占30%。长叶提灯藓LL1d的振幅占正常光周期的1/3, 而LL2d昼夜节律振幅无显著变化。脆枝青藓正常光周期时Pn在正午/午夜时达到最高和最低峰值, LL1d延续这一趋势, 但是LL2d峰值前移4 h。而Tr的峰值与Pn相比延后4 h (图1B), 同样受到昼夜节律调控。脆枝青藓的Tr在LL1d/LL2d条件下分别占正常光周期的29.6%和25.9% (r = 0.92); 大灰藓分别占正常光周期的58.6%和34.4%; 长叶提灯藓在LL1d振幅占正常光周期的56.0%。
图1
图1
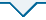
使用广义相加模型(GAM)拟合脆枝青藓、大灰藓和长叶提灯藓的净光合同化速率(Pn)、蒸腾速率(Tr)、非结构性碳水化合物和核心生物钟基因及节律调控相关基因昼夜节律。A、B中灰色部分对应于15 d正常光周期的最后一个夜晚。横坐标上的黑色和白色分别代表植物自然条件下的夜晚和白天, 灰色部分代表植物自然条件下黑暗但给予光照。C-I中灰色部分表示全光照条件下第二天(LL2d)外界条件下的夜晚。D-I分别为脆枝青藓生物钟调控相关基因的表达量。CDF1, CYCLING DOF FACTOR1; CKB, CASEIN KINASE II BETA CHAIN; CO, Constans; PAP1, PRODUCTION OF ANTHOCYANIN PIGMENT 1; PRR7, PSEUDO RESPONSE REGULATOR 7; PRR9, PSEUDO RESPONSE REGULATOR。
Fig. 1
Circadian rhythms of core circadian clock genes and rhythm regulation-related genes fitted to photosynthetic assimilation rate (Pn), transpiration rate (Tr), and non-structural carbon (NSC) in Brachythecium thraustum、Hypnum plumaeforme and Mnium lycopodioides using generalized additive model (GAM). The gray part in A and B corresponds to the last night of the 15 d normal photoperiod. The black and white parts on the horizontal coordinates represent the night and daytime of the plant under natural conditions, respectively, while the gray part represents the darkness of the plant under natural conditions but given light. The gray part in C-I represents the night under full light conditions on the second day (LL2d) under external conditions. D-I represent expression of clock-regulated genes in the moss Ceratodon purpureus. CDF1, CYCLING DOF FACTOR1; CKB, CASEIN KINASE II BETA CHAIN; CO, Constans; PAP1, PRODUCTION OF ANTHOCYANIN PIGMENT 1; PRR7, PSEUDO RESPONSE REGULATOR 7; PRR9, PSEUDO RESPONSE REGULATOR.
综上所述, 3种藓类植物光合作用均受到显著的昼夜节律调节, 其中脆枝青藓振幅最大、峰值位移最小, 因此以脆枝青藓为材料, 测量光合作用产物——NSC含量和相关基因昼夜间时序转录组。脆枝青藓LL2d的NSC含量变化呈现明显的节律性特征: 在白昼持续上升, 正午左右积累达到峰值; 夜间即使光照期间也持续下降, 午夜左右达到最低值, 与光合作用趋势相同; 相关性分析表明, NSC含量变化与Pn和Tr具备较强相关性(r = 0.94, p < 0.01)。值得关注的是: NSC中淀粉含量的日变化高于可溶性糖。
2.2 脆枝青藓在恒定光照下固碳耗水相关基因表达特点
2.2.1 生物钟基因的进化与节律表达
以拟南芥20个核心生物钟基因(CCA1、LHY、TOC1、PRR3、5、7、9、RVE1-8、LUX、ELF3/4, GI和ZTL)确定脆枝青藓的生物钟的直系同源基因及其表达特点。脆枝青藓具有4个核心生物钟基因家族的直系同源基因: 早间循环PRRs家族, 晚间循环ELF3/4、RVEs家族; 但缺少核心反馈环CCA1/LHY、TOC1的直系同源基因, 以及对TOC1等调节的GI、ZTL基因。如图1D、1E所示, 脆枝青藓核心生物钟基因PRRs在外界白天表达水平达到峰值, 具有显著的昼夜节律。而生物钟调控相关基因CYCLING DOF FACTOR 1 (CDF1)、Constans (CO)、PRODUCTION OF ANTHOCYANIN PIGMENT 1 (PAP1)和CASEIN KINASE II BETA CHAIN (CKB)同样具有显著的昼夜节律表达(图1F-1I)。
2.2.2 脆枝青藓气孔运动相关基因的进化与节律性表达
图2
图2
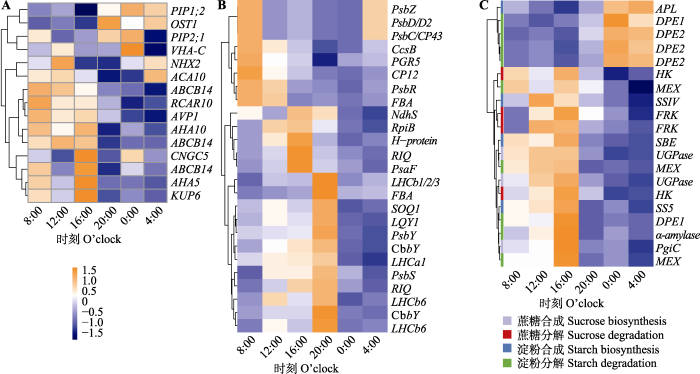
在恒定光照下脆枝青藓相关基因的表达。A, 15个显著的气孔相关节律性基因热图。B, 25个显著的光合相关节律性基因热图。C, 20个显著的非结构性碳水化合物相关节律性基因热图。热图采用了归一化表达量, 范围为-1.5到1.5。
Fig. 2
Expression of significant rhythmic genes related to stomata (A), photosynthesis(B) and non-structural carbohydrates (C) for Brachythecium thraustum under continuous illumination and constant environmental conditions. The heatmap used normalized expression levels, ranging from -1.5 to 1.5.
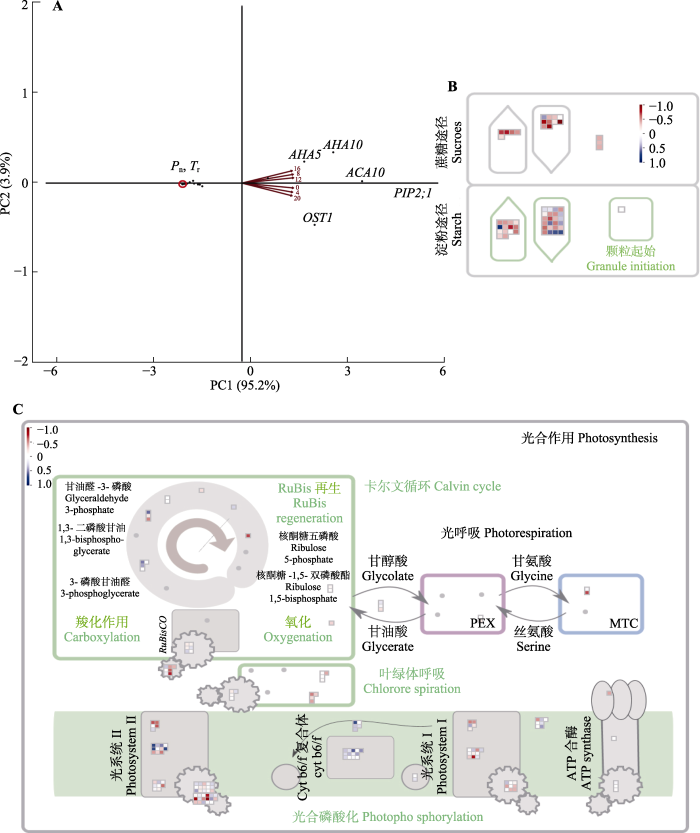
气孔运动为水分耗散等气体交换提供条件; 研究气体交换作用与气孔相关基因的相关性, 对Pn、Tr和脆枝青藓气孔运动相关基因表达进行主成分分析(图3A)。PC1概括95.2%的变量, 因此PC1能够较为充分地反应指标间关系。PC1负向端概括Pn、Tr以及RCAR10、ABCB14、KUP6等气孔调节相关基因的表达变化; 而PC1正向端概括PIP2;1、ACA10、AHA5、AVP1等胞内水分运输分配相关基因及OST1激酶的表达变化。
图3
图3
脆枝青藓相关基因的相关性分析。A, 15个显著的气孔相关节律性基因主成分(PC)分析图。B, 57个显著的碳固定相关节律性基因相关性图, 红色表示正相关, 蓝色表示负相关。C, 159个显著的光合相关节律性基因相关性图。热图采用了归一化表达量, 范围为-1到1。蓝色区域代表高表达量, 而红色区域则代表低表达量。ACA10, Ca2 +-ATP酶10; AHA5, 质膜H+-ATP酶; AHA10, H+-ATP酶10; MTC, PSII反应中心相关的膜内捕光复合物; OST1, 气孔开放因子1; PEX, 光系统II (PSII)外围蛋白质; PIP2;1, 水通道蛋白PIP2-1; Pn, 净光合速率; Tr, 蒸腾速率。
Fig. 3
Correlation analysis of genes related to Brachythecium thraustum. Principal Component (PC) analysis of 15 significant stomatal-related rhythmicity genes (A), 57 significant carbon fixation-related rhythmicity genes correlation plot (B) and 159 significant photosynthesis-related rhythmicity genes correlation plot (C) of Brachythecium thraustum. Red color indicates positive correlation, blue color indicates negative correlation. The heatmap used normalized expression levels, ranging from -1 to 1. In this scheme, blue regions represent high expression levels, while red regions represent low expression levels. ACA10, Ca2+-ATPase 10; AHA5, plasma membrane H+-ATPase 5; AHA10, H+-ATPase 10; MTC, membrane-intrinsic light-harvesting complexes associated with PSII reaction centers; OST1, open stomata 1; PEX, photosystem II (PSII) extrinsic proteins; PIP2;1, aquaporin PIP2-1; Pn, photosynthetic rate; Tr, transpiration rate.
2.2.3 脆枝青藓光合作用和NSC相关基因的进化与节律性表达
采用同样方法, 发现在脆枝青藓NSC相关的57个直系同源基因中, 其中与NSC含量相关的基因有20个(图2C), 占比35.1%; 15个基因的表达量在8:00上升, 在持续光照下于20:00下降, 与NSC含量变化趋势一致, 主要与淀粉合成、可溶性糖的合成与代谢相关; 剩余5个淀粉代谢相关基因(DPE1和DPE2)在夜间达到峰值, 与NSC含量变化呈负相关关系。
3 讨论
多数被子植物对光合作用具有节律响应, 蒸腾耗水作用缺乏节律性(Resco de Dios & Gessler, 2018); 然而本研究发现3种藓类植物均具有光合作用及蒸腾耗水节律。进一步采用脆枝青藓时序转录组分析, 发现超过一半的植物气孔运动相关的基因家族成员的表达具有较强节律性(图2A); 且这些基因表达与蒸腾耗水速率强相关, 表明蒸腾作用的节律性可以由气孔运动相关基因的节律调控进行解释。藓类植物是第一批脱离水生环境进入到陆地环境的陆生植物, 缺乏维管组织运输水分, 仅有1层细胞(极少情况下超过2层)的拟叶(即藓类植物的叶片)完成水分运输(Glime et al., 2007)。拟叶依靠气孔和细胞壁吸取水分并通过前者吸收CO2, 但缺少角质层保护, 因此对外界环境条件反应灵敏。除此前早已发现栖居环境中水分条件、光照对藓类的调节作用以外, 本研究表明藓类植物对于气孔周围组织的水分运输同样具有节律调节作用(图1B)。而且藓类植物气体交换指标(Pn和Tr)速率的节律调节振幅较强, 高于维管植物15%-25% (Resco de Dios & Gessler, 2018)的昼夜节律振幅。
植物气孔运动和蒸腾耗水作用由三方面决定: 一是叶片与大气之间VPD, 二是保卫细胞脱落酸(ABA)的调节, 三是水势变化(Chaves et al., 2008)。本研究中持续光照处理下VPD维持稳定, 排除其对节律调节的干扰。Ruszala等(2011)发现石松植物、卷柏(Selaginella moellendorffii)以及藓类Physcomitrella patens和Funaria hygrometrica中ABA诱导气孔关闭; 本研究基于藓类核心ABA信号通路(Hauser et al., 2011)成员的PYR/PYL/RCAR的节律变化发现ABA在藓类气体交换中起重要作用。更重要的是, 本研究表明藓类细胞的水势变化是气孔运动和蒸腾耗水节律调控的主要原因。尽管整体水势难以用压力室法测定, 但导致水势变化的保卫细胞水分/离子运输相关基因表达具有较强的节律, 间接证明了水势的主要作用: 如水分运输相关基因(PIP2;1、PIP1;2)、OST1激酶、保卫气孔离子通道(ABCB14、CNGC5、ACA10、 NHX2、KUP6)和提供质子动力势的ATP酶(AHA5、AHA10)、液泡H+-焦磷酸酶AVP1的节律变化。
迄今为止, 植物气体交换作用的昼夜节律机制尚未完全了解; 基于拟南芥的研究表明核心生物钟振荡器信号直接或者间接调控气体交换速率: 尤其光捕获、CO2吸收和气孔开放的节律(Jones, 2017)。进一步研究认为植物细胞中的生物钟通过分级振荡来调节气体交换作用的节律(Dodd et al., 2014; Endo, 2016)。人们普遍认为从藻类到被子植物CCA1是所有植物生物钟的基本组成部分。连同莱茵衣藻(Chlamydomonas reinhardtii)和光亮克里藻(Klebsormibium flaccidum)在内, 大多数植物生物钟理论模型都包含CCA1/LHY核心反馈环成分(Pokhilko et al., 2012; Fogelmark & Troein, 2014)。然而在本研究中, 脆枝青藓缺乏生物钟核心反馈环成分CCA1/LHY的同源物。Linde等(2017)的研究表明地钱(Marchantia polymorpha)和Anthoceros agrestis缺乏生物钟核心反馈环成分CCA1/LHY的同系物, 小立碗藓虽具有生物钟核心反馈环成分CCA1/LHY的同源物PpCCA1b, 但Okada等(2009)发现其在持续光照下PpCCA1b的生物钟核心功能失调。与小立碗藓一样, 脆枝青藓同样缺少GI、TOC1和ZTL同源物。在拟南芥中, GI和ZTL蛋白之间的相互作用调节TOC1的稳定性(Más et al., 2003; Kim et al., 2007)。但地钱中具有GI、TOC1和ZTL同源物, 表明这3个基因在藓类系谱分化早期丢失。GI是核心生物钟、糖类和光信号等不同信号通路因子和胁迫响元件; 在拟南芥等模式植物中发现GI通过促进CDF转录阻遏物, 激活关键的CO的表达; 脆枝青藓缺失GI, 因此CDF1完全依赖于PRRs, 其节律表达与PRRs相反, 进而CO与PRRs表达节律高度一致。由于藓类植物以配子体有性世代为主, 本研究对象为配子体, CO严格受PRRs调节能够保证其在有性世代受到节律严格调节, 而不会受到糖类等其他环境信号影响GI进而干扰CO变化。因此PRRs家族在脆枝青藓中承担了核心生物钟功能。
陆地植物生物钟的进化通常是核心生物钟基因数量的增加或形成更多的反馈环相互作用, 如从一个可以建模为简单双基因反馈环的绿藻生物钟进化到由许多基因形成的复杂调控网络的拟南芥生物钟。尽管如此, 由于藓类等低等植物类型对外界环境更为敏感, 因此其节律调节作用比被子植物等表现得更为明显; 此前对植物生物钟进化的研究(Ferrari et al., 2019)也表明这一点。
4 小结
综上所述, 藓类植物作为脱离水生环境进入到陆地环境的第一批陆生植物, 虽然缺少生物钟核心反馈环CCA1/LHY基因, 但依靠PRRs等生物钟基因仍然在光合/蒸腾作用、碳水化合物积累和相关基因表达上表现出较强的节律, 尤其在蒸腾耗水方面的节律调节作用超过被子植物。时序转录组研究表明藓类植物的蒸腾耗水节律作用主要由保卫细胞的水分/离子运输的节律性调节有关。本研究发现除环境水分、光照对藓类的调节作用以外, 生物节律对藓类植物的蒸腾耗水同样具有重要的调节作用; 为进一步研究陆生植物生物功能和生态作用的节律演化提供相关依据, 也为藓类植物的保育和利用提供了理论依据。
参考文献
Evolution of the stomatal regulation of plant water content
DOI:10.1104/pp.17.00078 PMID:28404725 [本文引用: 1]
Mutants of the biological clock in Chlamydomonas reinhardi
DOI:10.1093/genetics/70.4.537
PMID:5034771
[本文引用: 1]

A genetic analysis of the biological clock in Chlamydomonas reinhardi has been initiated. Of six wild-type strains tested (3 mt(+) and 3 mt(-)), five had periods close to 24 hr whereas one had a 21-hr period. Mutants with altered clock period have been isolated. The periods of 3 of these variant strains are temperature compensated. Genetic crosses involving a long-period mutant suggest that a single gene confers the long-period character, and in general clock-period length seems to be a useful phenotypic measure of alterations in the clock due to genetic differences. One phase mutant was found but its behavior was variable and the phase of the rhythm, relative to a light-dark transition which initiates the rhythm, does not seem to be reliable as a parameter of clock differences. No markers have yet been mapped.
Fast and sensitive protein alignment using DIAMOND
DOI:10.1038/nmeth.3176
PMID:25402007
[本文引用: 1]

The alignment of sequencing reads against a protein reference database is a major computational bottleneck in metagenomics and data-intensive evolutionary projects. Although recent tools offer improved performance over the gold standard BLASTX, they exhibit only a modest speedup or low sensitivity. We introduce DIAMOND, an open-source algorithm based on double indexing that is 20,000 times faster than BLASTX on short reads and has a similar degree of sensitivity.
Circadian rhythms of chemotaxis to ammonium and of methylammonium uptake in chlamydomonas
DOI:10.1104/pp.98.3.879
PMID:16668759
[本文引用: 1]

Chlamydomonas reinhardtii expresses a well-documented circadian rhythm of phototaxis, which peaks in the subjective daytime. We find that vegetative cells also express circadian rhythms of chemotaxis to ammonium and ammonium uptake (as gauged by uptake of [(14)C]methylammonium). The chemotaxis rhythm peaks in the subjective night. Methylammonium uptake is light dependent, and its rhythm peaks at subjective dawn. Unlike vegetative cells, gametes are not attracted to ammonium. We believe this to be the first report of a circadian rhythm of chemotaxis.
Photosynthesis under drought and salt stress: regulation mechanisms from whole plant to cell
DOI:10.1093/aob/mcn125 URL [本文引用: 1]
Molecular evolution of grass stomata
DOI:10.1016/j.tplants.2016.09.005 URL [本文引用: 1]
A cytosolic glucosyltransferase is required for conversion of starch to sucrose in Arabidopsis leaves at night
DOI:10.1111/tpj.2004.37.issue-6 URL [本文引用: 1]
The circadian regulation of photosynthesis
DOI:10.1007/s11120-013-9811-8
PMID:23529849
[本文引用: 3]

Correct circadian regulation increases plant productivity, and photosynthesis is circadian-regulated. Here, we discuss the regulatory basis for the circadian control of photosynthesis. We discuss candidate mechanisms underpinning circadian oscillations of light harvesting and consider how the circadian clock modulates CO2 fixation by Rubisco. We show that new techniques may provide a platform to better understand the signalling pathways that couple the circadian clock with the photosynthetic apparatus. Finally, we discuss how understanding circadian regulation in model systems is underpinning research into the impact of circadian regulation in crop species.
OrthoFinder: phylogenetic orthology inference for comparative genomics
DOI:10.1186/s13059-019-1832-y
PMID:31727128
[本文引用: 1]

Here, we present a major advance of the OrthoFinder method. This extends OrthoFinder's high accuracy orthogroup inference to provide phylogenetic inference of orthologs, rooted gene trees, gene duplication events, the rooted species tree, and comparative genomics statistics. Each output is benchmarked on appropriate real or simulated datasets, and where comparable methods exist, OrthoFinder is equivalent to or outperforms these methods. Furthermore, OrthoFinder is the most accurate ortholog inference method on the Quest for Orthologs benchmark test. Finally, OrthoFinder's comprehensive phylogenetic analysis is achieved with equivalent speed and scalability to the fastest, score-based heuristic methods. OrthoFinder is available at https://github.com/davidemms/OrthoFinder.
Tissue-specific circadian clocks in plants
DOI:10.1016/j.pbi.2015.11.003
PMID:26723003
[本文引用: 1]

Circadian clocks affect a large proportion of differentially expressed genes in many organisms. Tissue-specific hierarchies in circadian networks in mammals have been contentiously debated, whereas little attention has been devoted to the concept in plants, owing to technical difficulties. Recently, several studies have demonstrated tissue-specific circadian clocks and their coupling in plants, suggesting that plants possess a hierarchical network of circadian clocks. The following review summarizes recent studies describing the tissue-specific functions and properties of these circadian clocks and discusses the network structure and potential messengers that might share temporal information on such a network. Copyright © 2015 Elsevier Ltd. All rights reserved.
Kingdom-wide comparison reveals the evolution of diurnal gene expression in Archaeplastida
DOI:10.1038/s41467-019-08703-2
PMID:30760717
[本文引用: 1]

Plants have adapted to the diurnal light-dark cycle by establishing elaborate transcriptional programs that coordinate many metabolic, physiological, and developmental responses to the external environment. These transcriptional programs have been studied in only a few species, and their function and conservation across algae and plants is currently unknown. We performed a comparative transcriptome analysis of the diurnal cycle of nine members of Archaeplastida, and we observed that, despite large phylogenetic distances and dramatic differences in morphology and lifestyle, diurnal transcriptional programs of these organisms are similar. Expression of genes related to cell division and the majority of biological pathways depends on the time of day in unicellular algae but we did not observe such patterns at the tissue level in multicellular land plants. Hence, our study provides evidence for the universality of diurnal gene expression and elucidates its evolutionary history among different photosynthetic eukaryotes.
Rethinking transcriptional activation in the Arabidopsis circadian clock
DOI:10.1371/journal.pcbi.1003705 URL [本文引用: 1]
Chapter 2—Life cycles and morphology
// Glime JM.
Is the cell division cycle gated by a circadian clock? The case of Chlamydomonas reinhardtii
DOI:10.1083/jcb.129.4.1061
PMID:7744955
[本文引用: 1]

Circadian oscillators are known to regulate the timing of cell division in many organisms. In the case of Chlamydomonas reinhardtii, however, this conclusion has been challenged by several investigators. We have reexamined this issue and find that the division behavior of Chlamydomonas meets all the criteria for circadian rhythmicity: persistence of a cell division rhythm (a) with a period of approximately 24 h under free-running conditions, (b) that is temperature compensated, and (c) which can entrain to light/dark signals. In addition, a mutation that lengthens the circadian period of the phototactic rhythm similarly affects the cell division rhythm. We conclude that a circadian mechanism determines the timing of cell division in Chlamydomonas reinhardtii.
Full-length transcriptome assembly from RNA-Seq data without a reference genome
DOI:10.1038/nbt.1883
PMID:21572440
[本文引用: 1]

Massively parallel sequencing of cDNA has enabled deep and efficient probing of transcriptomes. Current approaches for transcript reconstruction from such data often rely on aligning reads to a reference genome, and are thus unsuitable for samples with a partial or missing reference genome. Here we present the Trinity method for de novo assembly of full-length transcripts and evaluate it on samples from fission yeast, mouse and whitefly, whose reference genome is not yet available. By efficiently constructing and analyzing sets of de Bruijn graphs, Trinity fully reconstructs a large fraction of transcripts, including alternatively spliced isoforms and transcripts from recently duplicated genes. Compared with other de novo transcriptome assemblers, Trinity recovers more full-length transcripts across a broad range of expression levels, with a sensitivity similar to methods that rely on genome alignments. Our approach provides a unified solution for transcriptome reconstruction in any sample, especially in the absence of a reference genome.
Integrating circadian dynamics with physiological processes in plants
DOI:10.1038/nrg3976
PMID:26370901
[本文引用: 2]

The plant circadian clock coordinates the responses to multiple and often simultaneous environmental challenges that the sessile plant cannot avoid. These responses must be integrated efficiently into dynamic metabolic and physiological networks essential for growth and reproduction. Many of the output pathways regulated by the circadian clock feed back to modulate clock function, leading to the appreciation of the clock as a central hub in a sophisticated regulatory network. In this Review, we discuss the circadian regulation of growth, flowering time, abiotic and biotic stress responses, and metabolism, as well as why temporal 'gating' of these processes is important to plant fitness.
Evolution of abscisic acid synthesis and signaling mechanisms
DOI:10.1016/j.cub.2011.03.015 URL [本文引用: 1]
Photosynthetic entrainment of the Arabidopsis thaliana circadian clock
DOI:10.1038/nature12603 [本文引用: 1]
Interplay of circadian rhythms and light in the regulation of photosynthesis-derived metabolism
ZEITLUPE is a circadian photoreceptor stabilized by GIGANTEA in blue light
DOI:10.1038/nature06132 [本文引用: 1]
RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome
DOI:10.1186/1471-2105-12-323
PMID:21816040
[本文引用: 1]

Background: RNA-Seq is revolutionizing the way transcript abundances are measured. A key challenge in transcript quantification from RNA-Seq data is the handling of reads that map to multiple genes or isoforms. This issue is particularly important for quantification with de novo transcriptome assemblies in the absence of sequenced genomes, as it is difficult to determine which transcripts are isoforms of the same gene. A second significant issue is the design of RNA-Seq experiments, in terms of the number of reads, read length, and whether reads come from one or both ends of cDNA fragments.;Results: We present RSEM, an user-friendly software package for quantifying gene and isoform abundances from single-end or paired-end RNA-Seq data. RSEM outputs abundance estimates, 95% credibility intervals, and visualization files and can also simulate RNA-Seq data. In contrast to other existing tools, the software does not require a reference genome. Thus, in combination with a de novo transcriptome assembler, RSEM enables accurate transcript quantification for species without sequenced genomes. On simulated and real data sets, RSEM has superior or comparable performance to quantification methods that rely on a reference genome. Taking advantage of RSEM's ability to effectively use ambiguously-mapping reads, we show that accurate gene-level abundance estimates are best obtained with large numbers of short single-end reads. On the other hand, estimates of the relative frequencies of isoforms within single genes may be improved through the use of paired-end reads, depending on the number of possible splice forms for each gene.;Conclusions: RSEM is an accurate and user-friendly software tool for quantifying transcript abundances from RNA-Seq data. As it does not rely on the existence of a reference genome, it is particularly useful for quantification with de novo transcriptome assemblies. In addition, RSEM has enabled valuable guidance for cost-efficient design of quantification experiments with RNA-Seq, which is currently relatively expensive.
Early evolution of the land plant circadian clock
DOI:10.1111/nph.2017.216.issue-2 URL [本文引用: 1]
Targeted degradation of TOC1 by ZTL modulates circadian function in Arabidopsis thaliana
DOI:10.1038/nature02163 [本文引用: 1]
Light-dependent regulation of cell division in Ostreococcus: evidence for a major transcriptional input
DOI:10.1104/pp.107.096149
PMID:17535824
[本文引用: 1]

Cell division often occurs at specific times of the day in animal and photosynthetic organisms. Studies in unicellular photosynthetic algae, such as Chlamydomonas or Euglena, have shown that the photoperiodic control of cell division is mediated through the circadian clock. However, the underlying mechanisms remain unknown. We have studied the molecular basis of light-dependent control of cell division in the unicellular green alga Ostreococcus. We found that cell division obeys a circadian oscillator in Ostreococcus. We provide evidence suggesting that the clock may, at least in part, regulate directly cell division independently of the metabolism. Combined microarray and quantitative real-time reverse transcription-polymerase chain reaction analysis of the main core cell cycle gene expression revealed an extensive transcriptional regulation of cell division by the photoperiod in Ostreococcus. Finally, transcription of the main core cell cycle genes, including cyclins and cyclin-dependent kinases, was shown to be under circadian control in Ostreococcus, suggesting that these genes are potential targets of the circadian clock in the control of cell division.
Circadian control of chloroplast transcription by a nuclear-encoded timing signal
DOI:10.1126/science.1230397
PMID:23493713
[本文引用: 2]

Circadian timekeeping in plants increases photosynthesis and productivity. There are circadian oscillations in the abundance of many chloroplast-encoded transcripts, but it is not known how the circadian clock regulates chloroplast transcription or the photosynthetic apparatus. We show that, in Arabidopsis, nuclear-encoded SIGMA FACTOR5 (SIG5) controls circadian rhythms of transcription of several chloroplast genes, revealing one pathway by which the nuclear-encoded circadian oscillator controls rhythms of chloroplast gene expression. We also show that SIG5 mediates the circadian gating of light input to a chloroplast-encoded gene. We have identified an evolutionarily conserved mechanism that communicates circadian timing information between organelles with distinct genetic systems and have established a new level of integration between eukaryotic circadian clocks and organelles of endosymbiotic origin.
Functional characterization of CCA1/LHY homolog genes, PpCCA1a and PpCCA1b, in the moss Physcomitrella patens
DOI:10.1111/j.1365-313X.2009.03979.x
PMID:19624471
[本文引用: 1]

The evolution of circadian clocks in land plants is not understood, because circadian rhythms have received little attention in plants other than angiosperms. We have characterized two genes, PpCCA1a and PpCCA1b, homologs of the Arabidopsis thaliana clock genes CCA1/LHY, from the moss Physcomitrella patens. PpCCA1a and PpCCA1b, together with angiosperm CCA1/LHY homologs, belong to the clock-associated single-myb gene family of green plants (including green algae and land plants). The accumulation of PpCCA1a and PpCCA1b mRNA showed rhythms with a period of approximately 1 day, phased as are those of angiosperm homologs, under 24 h light/dark cycles or in continuous dark. However, in marked contrast to angiosperm homologs, both genes showed arrhythmic profiles in continuous light. The timing of the PpCCA1b peak is determined by the time of the last light to dark transition, suggesting that the arrhythmicity in continuous light is due to dysfunction of the core clock. We generated single and double disruptants for PpCCA1a and PpCCA1b, and found that the double disruptants showed: (i) short periodicity and damped amplitude in the PpCCA1b rhythm, (ii) similar changes in the rhythmically expressed genes PpSIG5 and PpPRRa, and (iii) de-repression of PpCCA1b transcription levels, indicating negative feedback regulation. These observations indicate that the two genes are not merely structural homologs but also functional counterparts of CCA1/LHY. Together, our results illustrate similarities as well as divergence of the clock machineries between P. patens and A. thaliana, two distantly placed species in land plant phylogeny.
Differential involvement of the circadian clock in the expression of genes required for ribulose-1,5-bisphosphate carboxylase/oxygenase synthesis, assembly, and activation in Arabidopsis thaliana
We have investigated the role of the circadian clock in the regulation of expression of genes required for ribulose-1,5-bisphosphate carboxylase/oxygenase (Rubisco) synthesis, assembly, and activation. Circadian oscillations in RCA (the gene encoding Rubisco activase) and RBCS (the gene encoding Rubisco small subunit) mRNA accumulation, with peak abundance occurring soon after dawn, occur in Arabidopsis thaliana grown in a light-dark (LD) photoperiod. These oscillations persist in plants that have been transferred from LD to either continuous darkness (DD) or continuous light (LL). In contrast, CPN60[alpha] (the gene encoding [alpha]-chaperonin) and CPN60[beta] (the gene encoding [beta]-chaperonin) mRNA abundance oscillates in a diurnal, but not in a circadian, fashion. Although rapid damping of the circadian oscillation in RCA mRNA abundance is observed in Arabidopsis that have been grown in LD and then transferred to DD for 2 d, the circadian oscillations in RCA and RBCS mRNA abundance persist for at least five continuous cycles in LL, demonstrating the robustness of the circadian oscillator.
The clock gene circuit in Arabidopsis includes a repressilator with additional feedback loops
DOI:10.1038/msb.2012.6 URL [本文引用: 1]
Circadian rhythms have significant effects on leaf-to-canopy scale gas exchange under field conditions
DOI:10.1186/s13742-016-0149-y
PMID:27765071
[本文引用: 1]

Molecular clocks drive oscillations in leaf photosynthesis, stomatal conductance, and other cell and leaf-level processes over ~24 h under controlled laboratory conditions. The influence of such circadian regulation over whole-canopy fluxes remains uncertain; diurnal CO and HO vapor flux dynamics in the field are currently interpreted as resulting almost exclusively from direct physiological responses to variations in light, temperature and other environmental factors. We tested whether circadian regulation would affect plant and canopy gas exchange at the Montpellier European Ecotron. Canopy and leaf-level fluxes were constantly monitored under field-like environmental conditions, and under constant environmental conditions (no variation in temperature, radiation, or other environmental cues).We show direct experimental evidence at canopy scales of the circadian regulation of daytime gas exchange: 20-79 % of the daily variation range in CO and HO fluxes occurred under circadian entrainment in canopies of an annual herb (bean) and of a perennial shrub (cotton). We also observed that considering circadian regulation improved performance by 8-17 % in commonly used stomatal conductance models.Our results show that circadian controls affect diurnal CO and HO flux patterns in entire canopies in field-like conditions, and its consideration significantly improves model performance. Circadian controls act as a 'memory' of the past conditions experienced by the plant, which synchronizes metabolism across entire plant canopies.
Circadian regulation of photosynthesis and transpiration from genes to ecosystems
DOI:10.1016/j.envexpbot.2017.09.010 URL [本文引用: 3]
Soil phosphorous and endogenous rhythms exert a larger impact than CO2 or temperature on nocturnal stomatal conductance in Eucalyptus tereticornis
DOI:10.1093/treephys/tpt091 URL [本文引用: 1]
Land plants acquired active stomatal control early in their evolutionary history
DOI:10.1016/j.cub.2011.04.044
PMID:21658945
[本文引用: 1]

Stomata are pores that regulate plant gas exchange [1]. They evolved more than 400 million years ago [2, 3], but the origin of their active physiological responses to endogenous and environmental cues is unclear [2-6]. Recent research suggests that the stomata of lycophytes and ferns lack pore closure responses to abscisic acid (ABA) and CO(2). This evidence led to the hypothesis that a fundamental transition from passive to active control of plant water balance occurred after the divergence of ferns 360 million years ago [7, 8]. Here we show that stomatal responses of the lycophyte Selaginella [9] to ABA and CO(2) are directly comparable to those of the flowering plant Arabidopsis [10]. Furthermore, we show that the underlying intracellular signaling pathways responsible for stomatal aperture control are similar in both basal and modern vascular plant lineages. Our evidence challenges the hypothesis that acquisition of active stomatal control of plant carbon and water balance represents a critical turning point in land plant evolution [7, 8]. Instead, we suggest that the critical evolutionary development is represented by the innovation of stomata themselves and that physiologically active stomatal control originated at least as far back as the emergence of the lycophytes (circa 420 million years ago) [11].Copyright © 2011 Elsevier Ltd. All rights reserved.
MapMan4: a refined protein classification and annotation framework applicable to multi-omics data analysis
DOI:S1674-2052(19)30008-5
PMID:30639314
[本文引用: 1]

Genome sequences from over 200 plant species have already been published, with this number expected to increase rapidly due to advances in sequencing technologies. Once a new genome has been assembled and the genes identified, the functional annotation of their putative translational products, proteins, using ontologies is of key importance as it places the sequencing data in a biological context. Furthermore, to keep pace with rapid production of genome sequences, this functional annotation process must be fully automated. Here we present a redesigned and significantly enhanced MapMan4 framework, together with a revised version of the associated online Mercator annotation tool. Compared with the original MapMan, the new ontology has been expanded almost threefold and enforces stricter assignment rules. This framework was then incorporated into Mercator4, which has been upgraded to reflect current knowledge across the land plant group, providing protein annotations for all embryophytes with a comparably high quality. The annotation process has been optimized to allow a plant genome to be annotated in a matter of minutes. The output results continue to be compatible with the established MapMan desktop application.Copyright © 2019 The Authors. Published by Elsevier Inc. All rights reserved.