

植物生态学报 ›› 2009, Vol. 33 ›› Issue (1): 1-11.DOI: 10.3773/j.issn.1005-264x.2009.01.001
所属专题: 青藏高原植物生态学:种群生态学
• 研究论文 • 下一篇
程凯1,2, 孙坤3, 温红艳4, 张敏1, 贾东瑞2, 刘建全2,*( )
)
收稿日期:2007-12-06
接受日期:2008-03-04
出版日期:2009-12-06
发布日期:2009-01-30
通讯作者:
刘建全
作者简介:*E-mail: liujq@nwbip.ac.cn基金资助:
CHENG Kai1,2, SUN Kun3, WEN Hong-Yan4, ZHANG Min1, JIA Dong-Rui2, LIU Jian-Quan2,*( )
)
Received:2007-12-06
Accepted:2008-03-04
Online:2009-12-06
Published:2009-01-30
Contact:
LIU Jian-Quan
摘要:
研究具有邻近分布的近缘类群的遗传分化和谱系筛选对于进一步揭示物种形成机制具有重要意义。该文以沙棘属(Hippophae)临域分布的两个类群, 江孜沙棘(H. gyantsensis)和云南沙棘(H. rhamnoides subsp. yunnanensis)为研究对象, 进行群体水平上的母系分化研究。前一个种分布于西藏的中西部, 而后者主要分布于青藏高原东南部的云南西北部、四川西部和西藏东南部。两个类群在叶和果实性状存在明显区别。叶绿体在沙棘属植物中为母系遗传。共研究了两个类群14种群109个沙棘个体的叶绿体trnL-F、trnS-G序列; 序列排序后共发现11种单倍型, 江孜沙棘和云南沙棘分别有7种和6种, 两种单倍型为两个类群共享。分支分析和嵌套进化分析进一步表明, 两个类群之间的单倍型相互交错, 单倍型分化与形态上划分的两个类群不一致, 表明它们之间具有十分复杂的谱系筛选过程。这些发现明显不支持以前提出的有关江孜沙棘系统位置的假设。但是, 目前所获得的证据不能区分该物种究竟是通过异域分化还是同倍性杂交起源的。不同种群固定特有单倍型表明, 两个类群都在最后一次冰期可能存在多个避难所。
程凯, 孙坤, 温红艳, 张敏, 贾东瑞, 刘建全. 江孜沙棘和云南沙棘之间谱系分化和亲缘地理. 植物生态学报, 2009, 33(1): 1-11. DOI: 10.3773/j.issn.1005-264x.2009.01.001
CHENG Kai, SUN Kun, WEN Hong-Yan, ZHANG Min, JIA Dong-Rui, LIU Jian-Quan. MATERNAL DIVERGENCE AND PHYLOGEOGRAPHICAL RELATIONSHIPS BETWEEN HIPPOPHAE GYANTSENSIS AND H. RHAMNOIDES SUBSP. YUNNANENSIS. Chinese Journal of Plant Ecology, 2009, 33(1): 1-11. DOI: 10.3773/j.issn.1005-264x.2009.01.001
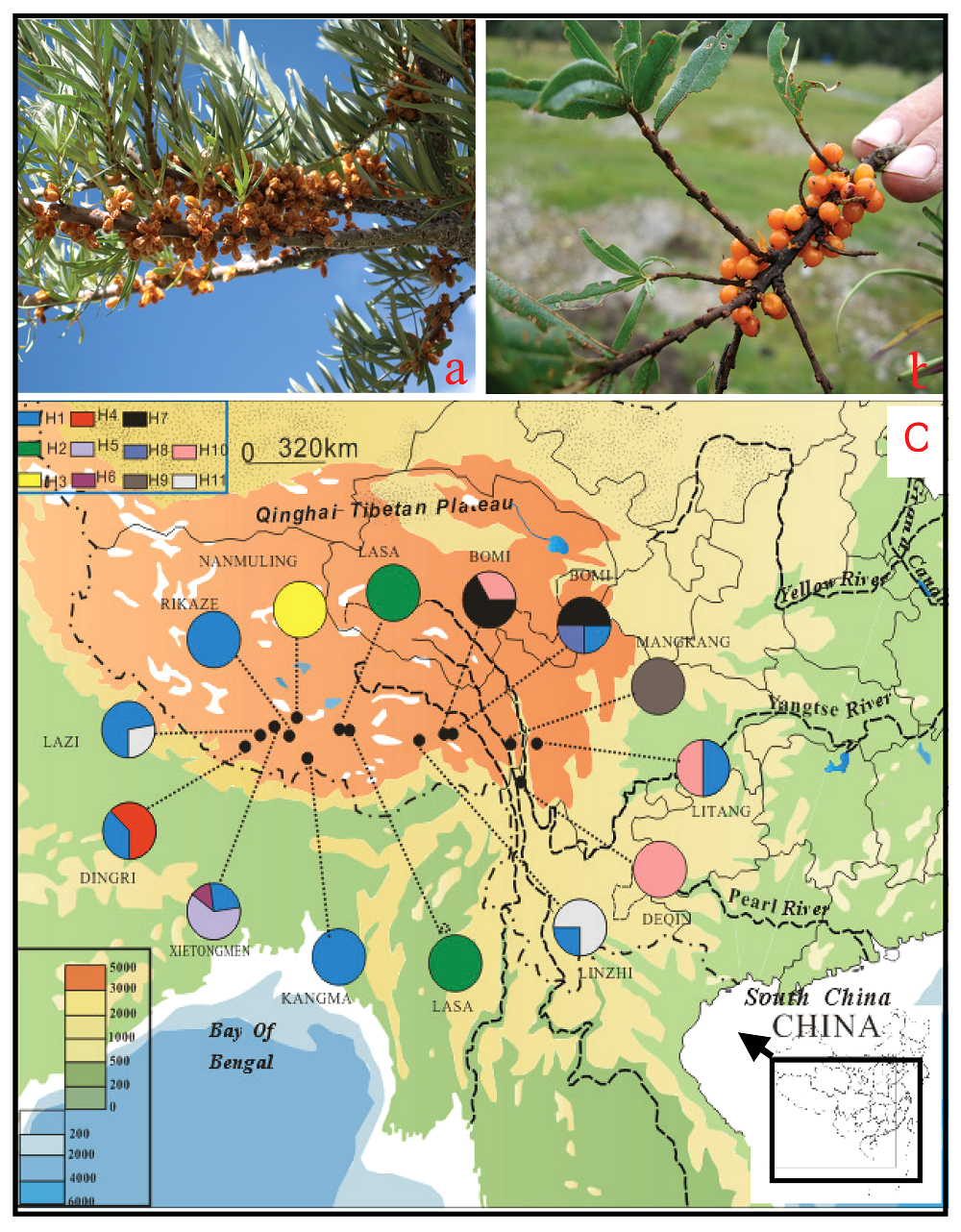
图1 江孜沙棘(南木林)(a)、云南沙棘(波密)(b)及两个类群(拉萨以西种群, 江孜沙棘; 林芝以东种群, 云南沙棘)取样种群的地理分布及相应的单倍型(c) H1~H11: 所获得的单倍型 Indicating the recouered haplotypes
Fig. 1 Hippophae gyantsensis (from Nanmulin)(a), H. rhamnoides subsp. yunnanensis (from Bomi) (b) and the sampled populations and the recovered haplotypes in each population of two taxa (Lasa westward, H. gyantsensis and Linzhi eastward, H. rhamnoides subsp. yunnanensis)(c)
| 种群号 Pop. No. | 采集号 Pop. code | 采样地 Location | 纬度 Latitude | 经度 Longitude | 海拔 Alt. (m) | 样本数 N | 遗传多样性指数 h | 核苷酸多样性指数 π |
|---|---|---|---|---|---|---|---|---|
| 1 | Liu2006252 | 西藏定日 Dingri, Tibet | 28o40′ | 087o11′ | 4 370 | 8 | 0.536 | 0.004 65 |
| 2 | Liu2006261 | 西藏拉孜 Lazi, Tibet | 29o12′ | 087o41′ | 3 970 | 9 | 0.389 | 0.001 76 |
| 3 | Liu2006262 | 西藏谢通门 Xietongmen, Tibet | 29o25′ | 088o13′ | 3 940 | 8 | 0.429 | 0.005 75 |
| 4 | Liu2006263 | 西藏日喀则 Rikazi, Tibet | 29o21′ | 088o49′ | 3 860 | 8 | 0.000 | 0.000 00 |
| 5 | Liu2006217 | 西藏康马 Kangma, Tibet | 28o48′ | 089o40′ | 4 150 | 9 | 0.000 | 0.000 00 |
| 6 | Liu2006227 | 西藏南木林 Nanmulin, Tibet | 29o41′ | 089o10′ | 4 030 | 7 | 0.000 | 0.000 00 |
| 7 | Liu2006271 | 西藏拉萨 Lasa, Tibet | 29o40′ | 091o06′ | 3 650 | 6 | 0.000 | 0.000 00 |
| 8 | Liu2006274 | 西藏拉萨 Lasa, Tibet | 29o36′ | 091o08′ | 3 730 | 8 | 0.000 | 0.000 00 |
| 9 | Liu2006286 | 西藏林芝 Linzhi, Tibet | 29o44′ | 094o43′ | 3 640 | 8 | 0.429 | 0.001 94 |
| 10 | Liu2006211 | 西藏波密 Bomi, Tibet | 29o12′ | 095o54′ | 3 860 | 6 | 0.533 | 0.005 31 |
| 11 | Liu2006287 | 西藏波密 Bomi, Tibet | 29o45′ | 095o58′ | 2 950 | 8 | 0.714 | 0.005 86 |
| 12 | Liu2006304 | 西藏芒康 Mangkang, Tibet | 29o42′ | 098o24′ | 3 430 | 9 | 0.000 | 0.000 00 |
| 13 | Liu2006309 | 云南德钦 Deqin, Yunnan | 28o27′ | 098o53′ | 3 470 | 7 | 0.000 | 0.000 00 |
| 14 | Liu2006317 | 四川理塘 Litang, Sichuan | 29o43′ | 100o24′ | 3 640 | 8 | 0.571 | 0.010 36 |
表1 沙棘材料来源(1~8, 江孜沙棘; 9~14, 云南沙棘)及叶绿体trnL-F与trnS-G单倍型在各种群中的单倍型多样性指数(h)及核苷酸多样性指数(π)
Table 1 Origin of materials, estimates of haplotype diversity (h) and nucleotide diversity (π) in the 14 populations 1-8: Hippophae gyantsensis; 9-14: H. rhamnoides subsp. yunnanensis h: Haplotype diversity π: Nucleotide diversity
| 种群号 Pop. No. | 采集号 Pop. code | 采样地 Location | 纬度 Latitude | 经度 Longitude | 海拔 Alt. (m) | 样本数 N | 遗传多样性指数 h | 核苷酸多样性指数 π |
|---|---|---|---|---|---|---|---|---|
| 1 | Liu2006252 | 西藏定日 Dingri, Tibet | 28o40′ | 087o11′ | 4 370 | 8 | 0.536 | 0.004 65 |
| 2 | Liu2006261 | 西藏拉孜 Lazi, Tibet | 29o12′ | 087o41′ | 3 970 | 9 | 0.389 | 0.001 76 |
| 3 | Liu2006262 | 西藏谢通门 Xietongmen, Tibet | 29o25′ | 088o13′ | 3 940 | 8 | 0.429 | 0.005 75 |
| 4 | Liu2006263 | 西藏日喀则 Rikazi, Tibet | 29o21′ | 088o49′ | 3 860 | 8 | 0.000 | 0.000 00 |
| 5 | Liu2006217 | 西藏康马 Kangma, Tibet | 28o48′ | 089o40′ | 4 150 | 9 | 0.000 | 0.000 00 |
| 6 | Liu2006227 | 西藏南木林 Nanmulin, Tibet | 29o41′ | 089o10′ | 4 030 | 7 | 0.000 | 0.000 00 |
| 7 | Liu2006271 | 西藏拉萨 Lasa, Tibet | 29o40′ | 091o06′ | 3 650 | 6 | 0.000 | 0.000 00 |
| 8 | Liu2006274 | 西藏拉萨 Lasa, Tibet | 29o36′ | 091o08′ | 3 730 | 8 | 0.000 | 0.000 00 |
| 9 | Liu2006286 | 西藏林芝 Linzhi, Tibet | 29o44′ | 094o43′ | 3 640 | 8 | 0.429 | 0.001 94 |
| 10 | Liu2006211 | 西藏波密 Bomi, Tibet | 29o12′ | 095o54′ | 3 860 | 6 | 0.533 | 0.005 31 |
| 11 | Liu2006287 | 西藏波密 Bomi, Tibet | 29o45′ | 095o58′ | 2 950 | 8 | 0.714 | 0.005 86 |
| 12 | Liu2006304 | 西藏芒康 Mangkang, Tibet | 29o42′ | 098o24′ | 3 430 | 9 | 0.000 | 0.000 00 |
| 13 | Liu2006309 | 云南德钦 Deqin, Yunnan | 28o27′ | 098o53′ | 3 470 | 7 | 0.000 | 0.000 00 |
| 14 | Liu2006317 | 四川理塘 Litang, Sichuan | 29o43′ | 100o24′ | 3 640 | 8 | 0.571 | 0.010 36 |
| 核苷酸位点Site 单倍型 Haplotypes | 5 5 | 5 6 | 2 0 0 | 2 4 2 | 5 1 4 | 5 1 6 | 5 2 2 | 5 2 6 | 5 4 4 | 5 6 9 | 5 8 1 | 6 0 4 | 6 0 5 | 6 1 0 | 6 1 4 | 6 4 9 | 6 8 1 | 6 8 6 | 6 9 5 | 6 9 7 | 7 0 2 | 7 5 9 | 7 7 7 | 7 8 0 | 7 8 1 | 8 1 3 | 8 4 9 | 8 8 3 | 9 0 4 | 9 2 6 | 9 3 6 | 9 4 7 | 9 7 1 | 9 7 4 | 9 7 5 | 1 0 0 1 | 1 0 0 8 | 1 0 1 4 | 1 0 1 5 | 1 0 3 2 | 1 0 7 3 | 1110 | 1124 | 1219 | 1224 | 1237 | 1416 | 1435 | 1483 | 1 4 98 | 1 5 0 9 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| H1 | A | T | T | G | C | T | C | T | A | T | T | A | A | T | C | A | A | C | A | T | A | T | A | T | G | T | G | C | T | T | C | △ | ● | A | ■ | ★ | ◇ | A | ㊣ | # | ⊙ | A | G | C | T | - | T | C | A | A | C |
| H2 | A | T | T | A | C | T | C | T | A | T | C | A | A | T | C | A | A | G | A | T | A | T | A | T | G | T | G | C | T | T | C | △ | ● | C | ■ | ★ | ◇ | A | ㊣ | - | ⊙ | A | G | A | T | - | T | C | C | A | C |
| H3 | T | G | C | G | C | T | C | C | C | T | C | A | A | T | C | C | A | G | G | G | C | G | G | T | G | G | G | C | T | T | C | △ | ● | A | ■ | ★ | ◇ | A | ㊣ | # | ⊙ | A | G | C | T | - | T | C | A | A | C |
| H4 | T | G | C | G | C | T | C | C | C | T | C | A | A | A | A | A | C | C | A | G | A | T | G | G | G | T | G | C | T | T | C | △ | ● | A | ■ | ★ | ◇ | A | ㊣ | # | ⊙ | A | G | C | T | - | T | C | A | A | C |
| H5 | T | G | C | G | T | C | T | C | C | A | C | T | C | T | C | A | C | C | A | G | A | T | G | G | A | T | A | C | T | T | C | △ | ● | A | ■ | ★ | ◇ | A | ㊣ | # | ⊙ | A | G | C | T | - | T | C | A | A | C |
| H6 | T | G | C | G | T | C | T | C | C | A | C | T | C | T | C | A | C | C | A | G | A | T | G | G | A | T | A | C | T | T | C | △ | ● | A | ■ | ★ | ◇ | A | ㊣ | - | ⊙ | A | G | C | T | - | T | C | A | A | C |
| H7 | T | G | C | G | C | T | C | C | C | A | C | A | A | A | A | A | C | C | A | G | A | T | G | G | G | T | G | C | T | A | C | - | - | A | - | - | - | - | - | - | - | G | G | A | G | * | T | C | C | - | C |
| H8 | T | G | C | G | C | T | C | C | C | T | C | A | A | A | A | A | C | C | A | G | A | T | G | G | G | T | G | C | T | T | C | - | - | A | - | - | - | - | - | - | - | G | G | A | G | * | T | C | C | - | C |
| H9 | T | G | C | G | C | T | C | C | C | T | C | A | A | T | C | C | A | G | G | G | C | G | G | T | G | G | G | T | C | T | G | - | ● | A | ■ | - | ◇ | C | ㊣ | - | ⊙ | G | A | A | T | * | T | G | C | - | T |
| H10 | T | G | C | G | T | C | T | C | C | A | C | T | C | T | C | A | C | C | A | G | A | T | G | G | A | T | A | C | T | A | C | - | ● | C | ■ | ★ | ◇ | A | ㊣ | - | ⊙ | G | G | A | T | * | G | C | C | - | C |
| H11 | A | T | T | G | C | T | C | C | A | T | T | A | A | T | C | A | A | C | A | T | A | T | A | T | G | T | G | C | T | A | C | - | ● | C | ■ | ★ | ◇ | A | ㊣ | - | ⊙ | G | G | A | T | * | G | C | C | - | C |
表2 沙棘属两个类群11种单倍型之间(trnL-F和 trnS-G区域)的序列区别(△●■★◇㊣#⊙*, 9个插入缺失)
Table 2 The sequence alignment of the 11 recovered haplotypes based trnL-F and trnS-G sequence in two taxa of Hippophae (△●■★◇㊣#⊙*, nine indels)
| 核苷酸位点Site 单倍型 Haplotypes | 5 5 | 5 6 | 2 0 0 | 2 4 2 | 5 1 4 | 5 1 6 | 5 2 2 | 5 2 6 | 5 4 4 | 5 6 9 | 5 8 1 | 6 0 4 | 6 0 5 | 6 1 0 | 6 1 4 | 6 4 9 | 6 8 1 | 6 8 6 | 6 9 5 | 6 9 7 | 7 0 2 | 7 5 9 | 7 7 7 | 7 8 0 | 7 8 1 | 8 1 3 | 8 4 9 | 8 8 3 | 9 0 4 | 9 2 6 | 9 3 6 | 9 4 7 | 9 7 1 | 9 7 4 | 9 7 5 | 1 0 0 1 | 1 0 0 8 | 1 0 1 4 | 1 0 1 5 | 1 0 3 2 | 1 0 7 3 | 1110 | 1124 | 1219 | 1224 | 1237 | 1416 | 1435 | 1483 | 1 4 98 | 1 5 0 9 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| H1 | A | T | T | G | C | T | C | T | A | T | T | A | A | T | C | A | A | C | A | T | A | T | A | T | G | T | G | C | T | T | C | △ | ● | A | ■ | ★ | ◇ | A | ㊣ | # | ⊙ | A | G | C | T | - | T | C | A | A | C |
| H2 | A | T | T | A | C | T | C | T | A | T | C | A | A | T | C | A | A | G | A | T | A | T | A | T | G | T | G | C | T | T | C | △ | ● | C | ■ | ★ | ◇ | A | ㊣ | - | ⊙ | A | G | A | T | - | T | C | C | A | C |
| H3 | T | G | C | G | C | T | C | C | C | T | C | A | A | T | C | C | A | G | G | G | C | G | G | T | G | G | G | C | T | T | C | △ | ● | A | ■ | ★ | ◇ | A | ㊣ | # | ⊙ | A | G | C | T | - | T | C | A | A | C |
| H4 | T | G | C | G | C | T | C | C | C | T | C | A | A | A | A | A | C | C | A | G | A | T | G | G | G | T | G | C | T | T | C | △ | ● | A | ■ | ★ | ◇ | A | ㊣ | # | ⊙ | A | G | C | T | - | T | C | A | A | C |
| H5 | T | G | C | G | T | C | T | C | C | A | C | T | C | T | C | A | C | C | A | G | A | T | G | G | A | T | A | C | T | T | C | △ | ● | A | ■ | ★ | ◇ | A | ㊣ | # | ⊙ | A | G | C | T | - | T | C | A | A | C |
| H6 | T | G | C | G | T | C | T | C | C | A | C | T | C | T | C | A | C | C | A | G | A | T | G | G | A | T | A | C | T | T | C | △ | ● | A | ■ | ★ | ◇ | A | ㊣ | - | ⊙ | A | G | C | T | - | T | C | A | A | C |
| H7 | T | G | C | G | C | T | C | C | C | A | C | A | A | A | A | A | C | C | A | G | A | T | G | G | G | T | G | C | T | A | C | - | - | A | - | - | - | - | - | - | - | G | G | A | G | * | T | C | C | - | C |
| H8 | T | G | C | G | C | T | C | C | C | T | C | A | A | A | A | A | C | C | A | G | A | T | G | G | G | T | G | C | T | T | C | - | - | A | - | - | - | - | - | - | - | G | G | A | G | * | T | C | C | - | C |
| H9 | T | G | C | G | C | T | C | C | C | T | C | A | A | T | C | C | A | G | G | G | C | G | G | T | G | G | G | T | C | T | G | - | ● | A | ■ | - | ◇ | C | ㊣ | - | ⊙ | G | A | A | T | * | T | G | C | - | T |
| H10 | T | G | C | G | T | C | T | C | C | A | C | T | C | T | C | A | C | C | A | G | A | T | G | G | A | T | A | C | T | A | C | - | ● | C | ■ | ★ | ◇ | A | ㊣ | - | ⊙ | G | G | A | T | * | G | C | C | - | C |
| H11 | A | T | T | G | C | T | C | C | A | T | T | A | A | T | C | A | A | C | A | T | A | T | A | T | G | T | G | C | T | A | C | - | ● | C | ■ | ★ | ◇ | A | ㊣ | - | ⊙ | G | G | A | T | * | G | C | C | - | C |
| 多样性参数 Diversity parameters | 合计 Total | 江孜沙棘 H. gyantsensis | 云南沙棘 H. rhamnoides subsp. yunnanensis |
|---|---|---|---|
| 种群 Populations | 15 | 8 | 6 |
| 单倍型 Haplotypes | 11 | 7 | 6 |
| HS | 0.270 (0.077 4) | 0.229(0.092 6) | 0.343 (0.147 1) |
| HT | 0.884 (0.043 8) | 0.850 (0.074 9) | 0.919 (0.065 5) |
| GST | 0.694 (0.090 2) | 0.730 (0.103 3) | 0.627 (0.169 9) |
| VS | 0.257 (0.079 6) | 0.166 (0.084 3) | 0.274 (0.131 6) |
| VT | 0.0934 (0.061 3) | 0.862 (0.126 0) | 1.036 (0.069 9) |
| NST | 0.725 (0.086 4) | 0.807 (0.108 0) | 0.735 (0.161 2) |
| NST -GST | 0.024 | 0.077 | 0.068 |
| h | 0.835 | 0.721 | 0.820 |
| π | 0.010 20 | 0.006 68 | 0.011 59 |
| Tajima’s D | 2.261 0* | 0..854 3 | 2.354 1* |
| Fu & Li’s D* | 2.083 8** | 1.883 7** | 1.908 6** |
表3 沙棘两个类群遗传多样性比较
Table 3 Genetic diversity and estimated indexes of two Hippophae taxa
| 多样性参数 Diversity parameters | 合计 Total | 江孜沙棘 H. gyantsensis | 云南沙棘 H. rhamnoides subsp. yunnanensis |
|---|---|---|---|
| 种群 Populations | 15 | 8 | 6 |
| 单倍型 Haplotypes | 11 | 7 | 6 |
| HS | 0.270 (0.077 4) | 0.229(0.092 6) | 0.343 (0.147 1) |
| HT | 0.884 (0.043 8) | 0.850 (0.074 9) | 0.919 (0.065 5) |
| GST | 0.694 (0.090 2) | 0.730 (0.103 3) | 0.627 (0.169 9) |
| VS | 0.257 (0.079 6) | 0.166 (0.084 3) | 0.274 (0.131 6) |
| VT | 0.0934 (0.061 3) | 0.862 (0.126 0) | 1.036 (0.069 9) |
| NST | 0.725 (0.086 4) | 0.807 (0.108 0) | 0.735 (0.161 2) |
| NST -GST | 0.024 | 0.077 | 0.068 |
| h | 0.835 | 0.721 | 0.820 |
| π | 0.010 20 | 0.006 68 | 0.011 59 |
| Tajima’s D | 2.261 0* | 0..854 3 | 2.354 1* |
| Fu & Li’s D* | 2.083 8** | 1.883 7** | 1.908 6** |
| 变异来源 Source of variation | 自由度 df | 方差 SS | 变异成分VC | 变异比例 Variation (%) | 固定指数 Fixation index | p |
|---|---|---|---|---|---|---|
| 类群间 Between taxa | 1 | 4.209 | 0.034 7 | 7.66 | FCT= 0.076 6 | 0.064 5 |
| 群体间 Among population within groups | 12 | 28.012 | 0.282 9 | 62.42 | FSC= 0.675 9 | < 0.001 |
| 群体内 Within populations | 95 | 12.889 | 0.135 7 | 29.92 | FST =0.700 8 | < 0.001 |
| 总计 Total | 108 | 45.110 | 0.453 4 |
表4 所有单倍型AMOVAs分析
Table 4 Analyses of molecular variance (AMOVAs) for all chlorotypes
| 变异来源 Source of variation | 自由度 df | 方差 SS | 变异成分VC | 变异比例 Variation (%) | 固定指数 Fixation index | p |
|---|---|---|---|---|---|---|
| 类群间 Between taxa | 1 | 4.209 | 0.034 7 | 7.66 | FCT= 0.076 6 | 0.064 5 |
| 群体间 Among population within groups | 12 | 28.012 | 0.282 9 | 62.42 | FSC= 0.675 9 | < 0.001 |
| 群体内 Within populations | 95 | 12.889 | 0.135 7 | 29.92 | FST =0.700 8 | < 0.001 |
| 总计 Total | 108 | 45.110 | 0.453 4 |
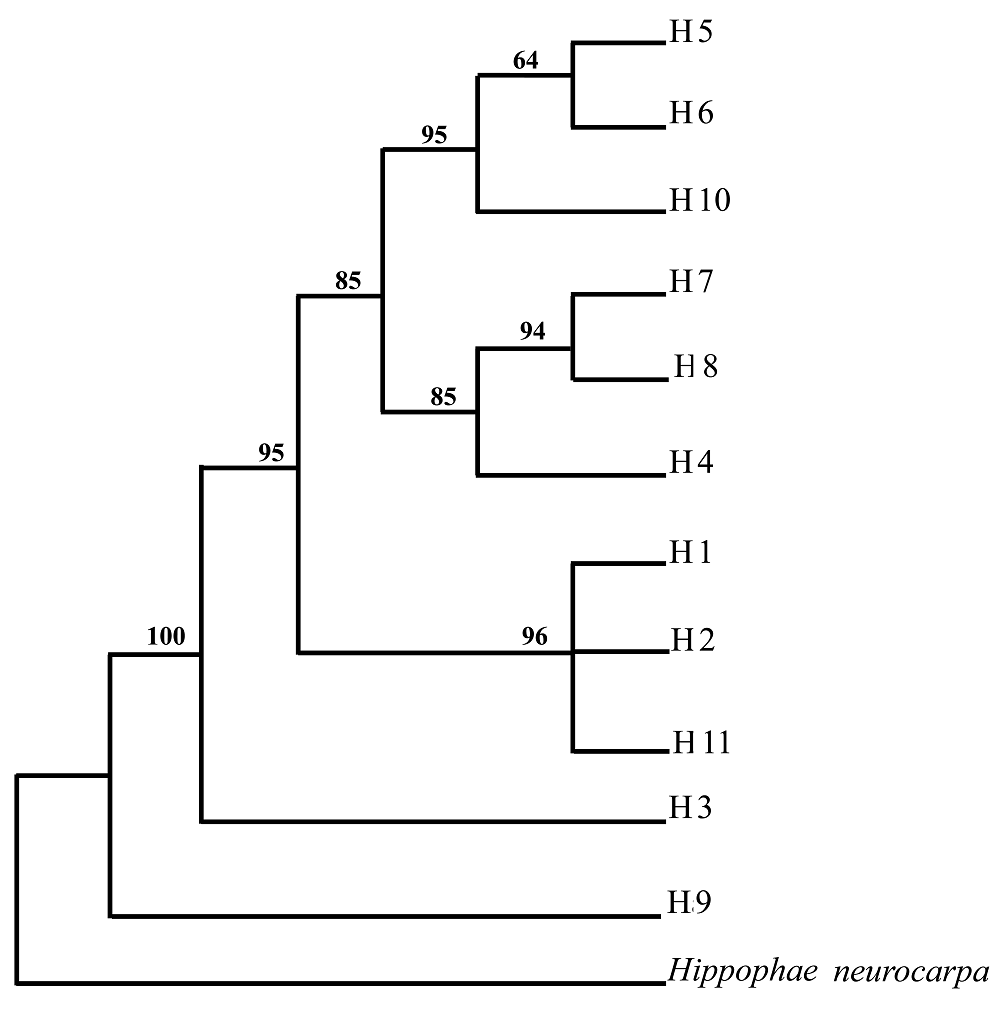
图2 基于两套叶绿体序列分析得到的唯一最简约树(只使用了突变信息位点) 分支下的数字为自展分析支持率(bootstrap值, 1 000次重复), 分支上的数字为枝长 Numbers below/above internodes are respectively bootstrap values based on 1 000 replicates and branch lengths H1~H11: 同图1 See Fig. 1
Fig. 2 The single maximum parsimonious (MP) tree based on two sets of cpDNA sequences (only mutations were used)
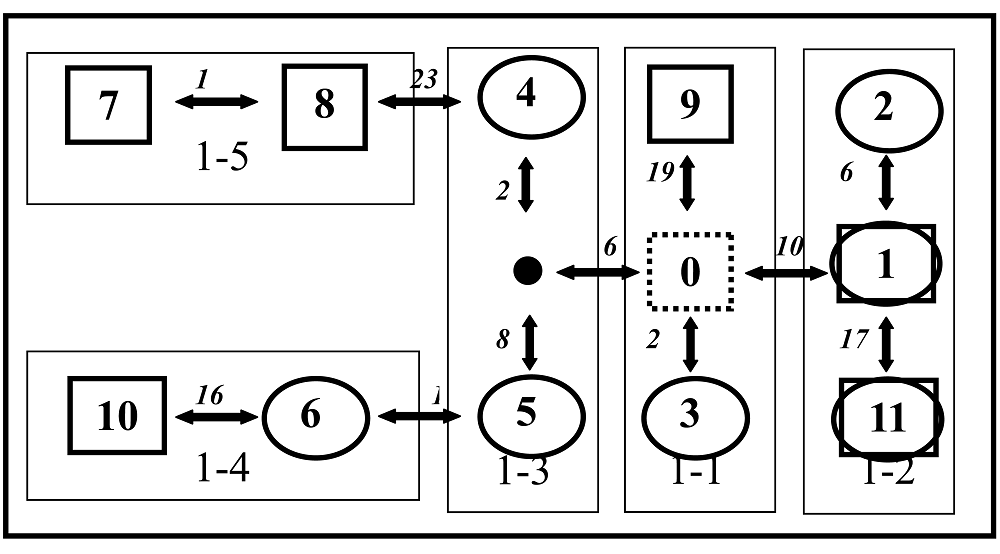
图3 沙棘两个类群11种叶绿体DNA单倍型的进化分支网络图 椭圆型和正方形分别表示在江孜沙棘和云南沙棘得到的单倍型, 斜体数字表示两节点间碱基差异步数 Elliptic and rectangle squares indicating the haplotypes respectively recovered in H. gyantsensis and H. rhamnoides subsp. yunnanensis
Fig. 3 The nested cladogram of cpDNA haplotypes in two taxa of Hippophae
| 分支 Clades | Chi平方统计 Chi-square statistic | p | 分支检索 Clade key | 推论 Inferences |
|---|---|---|---|---|
| 1-1 | 16.000 0 | <0.001 | 1-19-20-2-11-17 NO | 趋势不明显 Inconclusive outcome. |
| 1-2 | 90.592 9 | <0.001 | 1-2-3-5-15NO | 片段化或长距离扩张 Past fragmentation and/or long distance colonization |
| 1-3 | 10.000 0 | 0.006 | 1-19-20-2-11-17 NO | 趋势不明显 Inconclusive outcome |
| 1-4 | 14.000 0 | 0.075 | 1-19-20-2-11-17 NO | 趋势不明显 Inconclusive outcome |
| 1-5 | 1.666 7 | 0.479 | 1-2-11-17 NO | 趋势不明显 Inconclusive outcome |
| 整个分枝 Total cladogram | 318.775 5 | <0.001 | 1-2-11-17 NO | 趋势不明显 Inconclusive outcome |
表6 根据嵌套分支分析检索得出的结论
Table 6 The inference from the nested clade analysis following the given keys
| 分支 Clades | Chi平方统计 Chi-square statistic | p | 分支检索 Clade key | 推论 Inferences |
|---|---|---|---|---|
| 1-1 | 16.000 0 | <0.001 | 1-19-20-2-11-17 NO | 趋势不明显 Inconclusive outcome. |
| 1-2 | 90.592 9 | <0.001 | 1-2-3-5-15NO | 片段化或长距离扩张 Past fragmentation and/or long distance colonization |
| 1-3 | 10.000 0 | 0.006 | 1-19-20-2-11-17 NO | 趋势不明显 Inconclusive outcome |
| 1-4 | 14.000 0 | 0.075 | 1-19-20-2-11-17 NO | 趋势不明显 Inconclusive outcome |
| 1-5 | 1.666 7 | 0.479 | 1-2-11-17 NO | 趋势不明显 Inconclusive outcome |
| 整个分枝 Total cladogram | 318.775 5 | <0.001 | 1-2-11-17 NO | 趋势不明显 Inconclusive outcome |
| 零步 Zero-step | 一步 One-step | 两步 Two-step | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 单倍型 Haplotype | P | Dc | Dn | 分枝Clades | P | Dc | Dn | 分枝Clades | P | Dc | Dn | ||
| H11 | T | 4.308 S | 62.946S | 1-1 | T | 114.035 S | 114.462 S | 2-1 | T | 318.000 S | 333.484 S | ||
| H2 | T | 240.918 L | 310.847 L | ||||||||||
| H1 | I | 362.262 | 378.008 | 1-2 | I | 362.262 | 378.008 L | ||||||
| H9 | T | 0.000 S | 445.873 | 1-3 | T | 445.898 L | 447.296 L | ||||||
| H3 | T | 0.000 S | 445.923 L | ||||||||||
| I-T | - | 0.000 L | 0.050 L | ||||||||||
| I-T | 55.127 | 106.036 L | |||||||||||
| H4 | I | 0.000 S | 74.645 L | 1-4 | I | 63.913 S | 494.868 | 2-2 | T | 485.997 L | 480.024 L | ||
| H5 | I | 0.000 S | 55.864 S | ||||||||||
| H6 | I | 0.000 | 976.800 | 1-5 | T | 396.468 | 580.510 L | ||||||
| H10 | T | 91.000 | 127.444 | ||||||||||
| I-T | - | -91.000 | 849.356 | ||||||||||
| H7 | T | 0.000 | 108.520 | 1-6 | T | 341.659 | 343.437 S | ||||||
| H8 | I | 272.171 | 236.337 | ||||||||||
| I-T | - | 272.191 | 127.818 | ||||||||||
| I-T | -309.718 S | 13.138 | |||||||||||
表5 沙棘两个类群单倍型嵌套分支分析
Table 5 Nested clade analyses of the recovered haplotypes
| 零步 Zero-step | 一步 One-step | 两步 Two-step | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 单倍型 Haplotype | P | Dc | Dn | 分枝Clades | P | Dc | Dn | 分枝Clades | P | Dc | Dn | ||
| H11 | T | 4.308 S | 62.946S | 1-1 | T | 114.035 S | 114.462 S | 2-1 | T | 318.000 S | 333.484 S | ||
| H2 | T | 240.918 L | 310.847 L | ||||||||||
| H1 | I | 362.262 | 378.008 | 1-2 | I | 362.262 | 378.008 L | ||||||
| H9 | T | 0.000 S | 445.873 | 1-3 | T | 445.898 L | 447.296 L | ||||||
| H3 | T | 0.000 S | 445.923 L | ||||||||||
| I-T | - | 0.000 L | 0.050 L | ||||||||||
| I-T | 55.127 | 106.036 L | |||||||||||
| H4 | I | 0.000 S | 74.645 L | 1-4 | I | 63.913 S | 494.868 | 2-2 | T | 485.997 L | 480.024 L | ||
| H5 | I | 0.000 S | 55.864 S | ||||||||||
| H6 | I | 0.000 | 976.800 | 1-5 | T | 396.468 | 580.510 L | ||||||
| H10 | T | 91.000 | 127.444 | ||||||||||
| I-T | - | -91.000 | 849.356 | ||||||||||
| H7 | T | 0.000 | 108.520 | 1-6 | T | 341.659 | 343.437 S | ||||||
| H8 | I | 272.171 | 236.337 | ||||||||||
| I-T | - | 272.191 | 127.818 | ||||||||||
| I-T | -309.718 S | 13.138 | |||||||||||
| [1] | Avise JC (2000). Phylogeography: The History and Formation of Species. Harvard University Press, London. |
| [2] | Bartish IV, Jeppsson N, Bartish GI, Lu R, Nybom H (2000). Inter- and intraspecific genetic variation in Hippophae(Elaeagnaceae) investigated by RAPD markers. Plant Systematics and Evolution, 225, 81-101. |
| [3] | Bartish IV, Jeppsson N, Nybom H, Swenson U (2002). Phylogeny of Hippophae (Elaeagnaceae) inferred from parsimony analysis of chloroplast DNA and morphology. Systematic Botany, 27, 41-54. |
| [4] |
Bartish IV, Kadereit JW, Comes HP (2006). Late Quaternary history of Hippophae rhamnoides L. (Elaeagnaceae) inferred from chalcone synthase intron (Chsi) sequences and chloroplast DNA variation. Molecular Ecology, 15, 897-915.
DOI URL PMID |
| [5] |
Colosimo PF, Hosemann KE, Balabhadra S, Villarreal GJ, Dickson M, Grimwood J, Schmutz J, Myers RM, Schluter D, Kingsley DM (2005). Widespread parallel evolution in sticklebacks by repeated fixation of ectodysplasin alleles. Science, 307, 1928-1933.
DOI URL PMID |
| [6] | Doyle JJ, Doyle JL (1987). A rapid DNA isolation procedure for small quantities of fresh leaf material. Phytochemical Bulletin, 19, 11-15. |
| [7] |
Excoffier L, Laval G, Schneider S (2005). ARLEQUIN (version 3.0): an integrated software package for population genetics data analysis. Evolutionary Bioinformatics, 1, 47-50.
URL PMID |
| [8] |
Fu YX (1997). Statistical tests of neutrality of mutations against population growth, hitchhiking and background selection. Genetics, 147, 915-925.
URL PMID |
| [9] |
Hewitt G (2000). The genetic legacy of the Quaternary ice ages. Nature, 405, 907-913.
DOI URL PMID |
| [10] |
Hyvonen J (1996). On phylogeny of Hippophae(Elaeagnaceae). Nordic Journal of Botany, 16, 51-62.
DOI URL |
| [11] | Lian YS (廉永善) (2000). The Plant Biology and Chemistry of Hippophae (沙棘属植物生物学和化学). Gansu Science and Technology Press, Lanzhou. (in Chinese) |
| [12] | Lian YS, Chen XL, Lian H (1998). Systematic classification of the genus Hippophae L. Seabuckthorn Research, 1, 13-23. |
| [13] |
Meng LH (孟丽华), Yang HL (杨惠玲), Wu GL (吴桂莉), Wang YJ (王玉金) (2008). Phylogeography of Hippophae neurocarpa (Elaeagnaceae) inferred from the chloroplast DNA trnL-F sequence variation. Journal of Systematics and Evolution (植物分类学报), 46, 32-40. (in Chinese with English abstract)
DOI URL |
| [14] |
Newton AC, Allnot TR, Gillies ACM, Lowe AJ, Ennos RA (1999). Molecular phylogeography intraspecific variation and conservation of tree species. Trends in Ecology and Evolution, 14, 140-145.
DOI URL PMID |
| [15] |
Petit RJ, Aguinagalde I, de Beaulieu JL, Bittkau C, Brewer S, Cheddadi R, Ennos R, Fineschi S, Grivet D, Lascoux M, Mohanty A, Muller-Starck GM, Demesure-Musch B, Palme A, Martin JP, Rendell S, Vendramin GG (2003). Glacial refugia: hotspots but not melting pots of genetic diversity. Science, 300, 1563-1565.
DOI URL PMID |
| [16] |
Pons O, Petit RJ (1995). Estimation, variance and optimal sampling of gene diversity. I. Haploid locus. Theoretical and Applied Genetics, 90, 462-470.
DOI URL PMID |
| [17] |
Posada D, Crandall KA, Templeton AR (2000). GEODIS: a program for the cladistic nested analysis of the geographical distribution of genetic haplotypes. Molecular Ecology, 9, 487-488.
DOI URL PMID |
| [18] |
Rieseberg LH (1997). Hybrid origins of plant species. Annual Review of Ecology and Systematics, 28, 359-389.
DOI URL |
| [19] |
Rieseberg LH, Raymond O, Rosenthal DM, Lai Z, Livingstone K, Nakazato T, Durphy JL, Schwarzbach AE, Donovan LA, Lexer C (2003). Major ecological transitions in annual sunflowers facilitated by hybridization. Science, 301, 1211-1216.
DOI URL PMID |
| [20] |
Rieseberg LH, Widmer A, Arntz MA, Burke JM (2002). Directional selection is the primary cause of phenotypic diversification. Proceedings of the National Academy of Sciences of the United States of America, 99, 12242-12245.
DOI URL PMID |
| [21] |
Rieseberg LH, Wood TE, Baack E (2006). The nature of plant species. Nature, 440, 524-527.
DOI URL PMID |
| [22] | Rousi A (1971). The genus Hippophae L. A taxonomic study. Annales Botanici Fennici, 8, 177-227. |
| [23] |
Rozas J, Sanchez-Delbarrio JC, Messeguer X, Rozas R (2003). DnaSP, DNA polymorphism analyses by the coalescent and other methods. Bioinformatics, 19, 2496-2497.
DOI URL PMID |
| [24] | Shi YF (施雅风), Li JJ (李吉均), Li BY (李炳元) (1998). Uplift and Environmental Changes of Qinghai-Tibetan Plateau in the Late Cenozoic (青藏高原晚新生代隆升与环境变化). Guangdong Science and Technology Press, Guangzhou. (in Chinse) |
| [25] |
Song BH, Wang XQ, Wang XR, Ding KY, Hong DY (2003). Cytoplasmic composition in Pinus densata and population establishment of the diploid hybrid pine. Molecular Ecology, 12, 2995-3001.
DOI URL PMID |
| [26] |
Sun K, Chen XL, Ma RJ, Li CB, Wang Q, Ge S (2002). Molecular phylogenetics of Hippophae L.(Elaeagnaceae) based on the internal transcribed spacer (ITS) sequences of nrDNA. Plant Systematics and Evolution, 235, 121-134.
DOI URL |
| [27] | Sun K, Ma RJ, Chen XL, Li CB, Ge S (2003). Hybrid origin of the diploid species Hippophae goniocarpa evidenced by the internal transcribed spacers (ITS) of nuclear rDNA. Belgian Journal of Botany, 136, 91-96. |
| [28] | Swofford DL (2000). PAUP*. Phylogenetic Analysis Using Parsimony (*and Other Methods). Version 4. Sinauer Associates. Sunderland, Massachusetts. |
| [29] |
Taberlet P, Gielly L, Pautou G, Bouvet J (1991). Universal primers for amplification of three non-coding regions of chloroplast DNA. Plant Molecular Biology, 17, 1105-1109.
DOI URL PMID |
| [30] |
Templeton AR, Crandall KA, Sing CF (1992). A cladistic analysis of phenotypic associations with haplotypes inferred from restriction endonuclease mapping and DNA sequence data. III. Cladogram estimation. Genetics, 132, 619-633.
URL PMID |
| [31] |
Thompson JD, Higgins DG, Gibson TJ (1994). CLUST- ALW: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Research, 22, 4673-4680.
DOI URL PMID |
| [32] |
Wang AL, Schluetz F, Liu JQ (2008). Molecular evidence for double maternal origins of the diploid hybrid Hippophae goniocarpa(Elaeagnaceae). Botanical Journal of the Linnean Society, 156, 111-118.
DOI URL |
| [33] |
Wang XR, Szmidt AE, Savolainen O (2001). Genetic composition and diploid hybrid speciation of a high mountain pine, Pinus densata, native to the Tibetan Plateau. Genetics, 159, 337-346.
URL PMID |
| [34] | Weir BS (1996). Genetic Data Analysis II. Sinauer & Associates, Sunderland, Massachusetts. |
| [35] |
Whibley AC, Langlade NB, Andalo C, Hanna AI, Bangham A, Thebaud C, Coen E (2006). Evolutionary paths underlying flower color variation in Antirrhinum. Science, 313, 963-967.
DOI URL PMID |
| [36] |
Zhang Q (张茜), Yang R (杨锐), Wang Q (王钦), Liu JQ (刘建全) (2005). Phylogeography of Juniperus przewalskii (Cupressaceae) inferred from the chloroplast DNA trnT-trnF sequence variation. Acta Phytotaxonomica Sinica (植物分类学报), 43, 503-512. (in Chinese with English abstract)
DOI URL |
| [1] | 范紫腾, 毋钰灵, 王新菊, 李太强, 高江云. 共生真菌对兰科植物种间杂交后代种子萌发的效应[J]. 植物生态学报, 2019, 43(4): 374-382. |
| [2] | 张勃, 孙杉, 张志强, 李庆军. 杠杆状雄蕊及其进化生态学意义[J]. 植物生态学报, 2010, 34(1): 89-99. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||
Copyright © 2022 版权所有 《植物生态学报》编辑部
地址: 北京香山南辛村20号, 邮编: 100093
Tel.: 010-62836134, 62836138; Fax: 010-82599431; E-mail: apes@ibcas.ac.cn, cjpe@ibcas.ac.cn
备案号: 京ICP备16067583号-19