

Chin J Plant Ecol ›› 2025, Vol. 49 ›› Issue (10): 1626-1642.DOI: 10.17521/cjpe.2024.0445 cstr: 32100.14.cjpe.2024.0445
Special Issue: 濒危植物种群特征与保护
• Research Articles • Previous Articles Next Articles
LU Zi-Jia1,2, WANG Tian-Rui1, ZHENG Si-Si1, MENG Hong-Hu3,4, CAO Jian-Guo2, Gregor KOZLOWSKI1,5,6, SONG Yi-Gang1,*( )
)
Received:2024-12-09
Accepted:2025-02-07
Online:2025-10-20
Published:2025-11-20
Contact:
SONG Yi-Gang
Supported by:LU Zi-Jia, WANG Tian-Rui, ZHENG Si-Si, MENG Hong-Hu, CAO Jian-Guo, Gregor KOZLOWSKI, SONG Yi-Gang. Environmental adaptive genetic variation and genetic vulnerability of relict plant Pterocarya hupehensis[J]. Chin J Plant Ecol, 2025, 49(10): 1626-1642.
Add to citation manager EndNote|Ris|BibTeX
URL: https://www.plant-ecology.com/EN/10.17521/cjpe.2024.0445
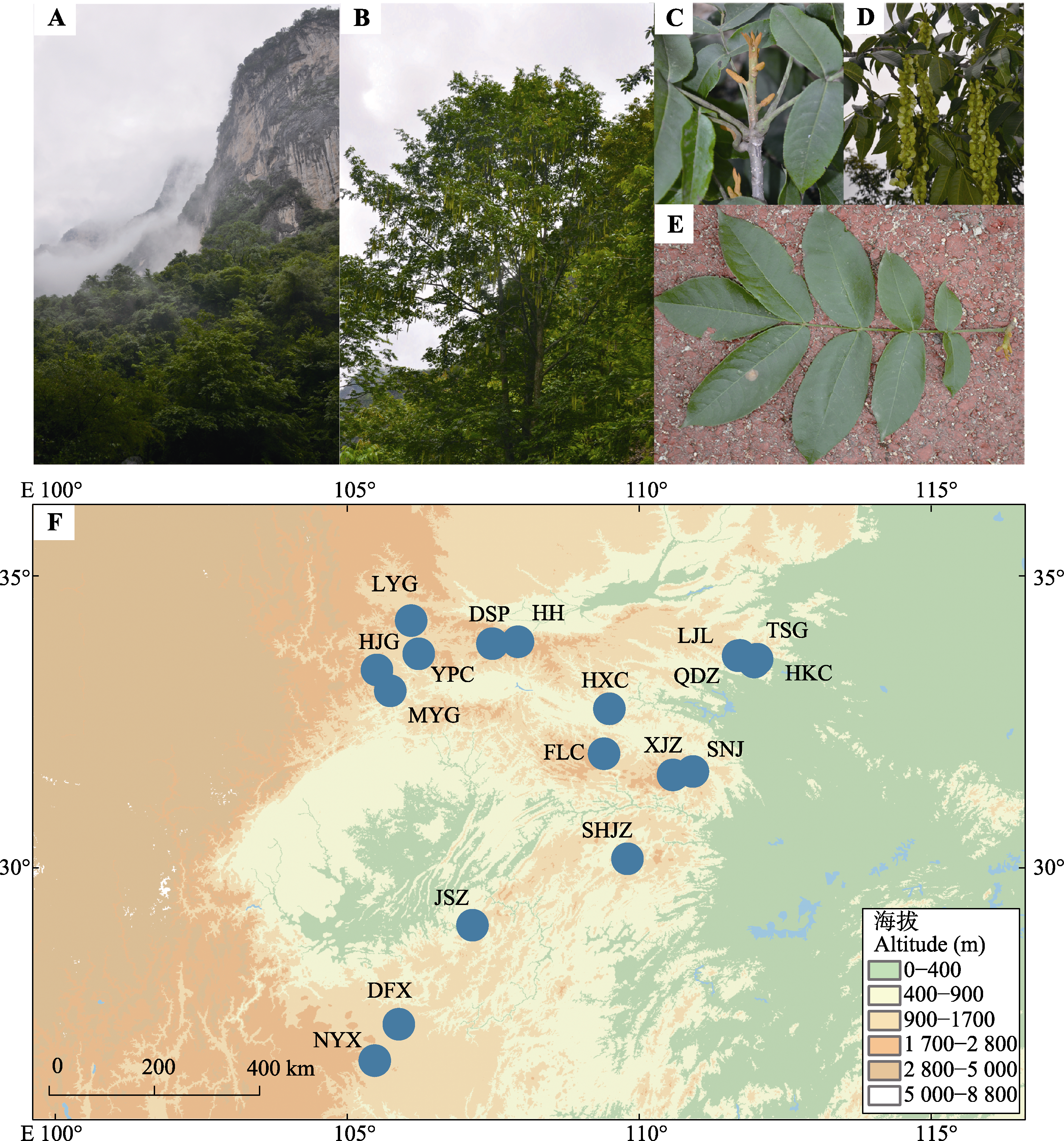
Fig. 1 Habitat of Pterocarya hupehensis (A), plant morphology (B-E), and collection sites of P. hupehensis (F). The information of collection sites is shown in Table 1.
| 种群编号 Code | 采样点 Site | 经度 Longitude (° E) | 纬度 Latitude (° N) | 个体数 n |
|---|---|---|---|---|
| HH | 陕西西安 Xi’an, Shaanxi | 107.93 | 33.87 | 6 |
| HXC | 陕西安康 Ankang, Shaanxi | 109.49 | 32.72 | 3 |
| FLC | 陕西安康 Ankang, Shaanxi | 109.40 | 31.95 | 5 |
| DSP | 陕西宝鸡 Baoji, Shaanxi | 107.49 | 33.84 | 4 |
| QDZ | 河南南阳 Nanyang, Henan | 111.97 | 33.52 | 6 |
| HKC | 河南南阳 Nanyang, Henan | 112.02 | 33.57 | 6 |
| TSG | 河南南阳 Nanyang, Henan | 111.72 | 33.63 | 6 |
| LJL | 河南南阳 Nanyang, Henan | 111.70 | 33.63 | 4 |
| LYG | 甘肃天水 Tianshui, Gansu | 106.10 | 34.23 | 5 |
| YPC | 甘肃陇南 Longnan, Gansu | 106.23 | 33.67 | 5 |
| HJG | 甘肃康县 Kang Xian, Gansu | 105.51 | 33.39 | 5 |
| MYG | 甘肃阳坝镇 Yangba, Gansu | 105.74 | 33.03 | 6 |
| SNJ | 湖北神农架 Shennongjia, Hubei | 110.92 | 31.65 | 7 |
| XJZ | 湖北神农架 Shennongjia, Hubei | 110.58 | 31.59 | 11 |
| SHJZ | 湖北恩施 Enshi, Hubei | 109.80 | 30.16 | 11 |
| JSZ | 重庆金山镇 Jinshan, Chongqing | 107.14 | 29.02 | 7 |
| DFX | 贵州毕节 Bijie, Guizhou | 105.88 | 27.33 | 12 |
| NYX | 贵州毕节 Bijie, Guizhou | 105.47 | 26.70 | 13 |
Table 1 Sample coding, sampling locations, and individual numbers of 18 Pterocarya hupehensis populations based on restriction-site associated DNA-sequnencing (RAD-seq)
| 种群编号 Code | 采样点 Site | 经度 Longitude (° E) | 纬度 Latitude (° N) | 个体数 n |
|---|---|---|---|---|
| HH | 陕西西安 Xi’an, Shaanxi | 107.93 | 33.87 | 6 |
| HXC | 陕西安康 Ankang, Shaanxi | 109.49 | 32.72 | 3 |
| FLC | 陕西安康 Ankang, Shaanxi | 109.40 | 31.95 | 5 |
| DSP | 陕西宝鸡 Baoji, Shaanxi | 107.49 | 33.84 | 4 |
| QDZ | 河南南阳 Nanyang, Henan | 111.97 | 33.52 | 6 |
| HKC | 河南南阳 Nanyang, Henan | 112.02 | 33.57 | 6 |
| TSG | 河南南阳 Nanyang, Henan | 111.72 | 33.63 | 6 |
| LJL | 河南南阳 Nanyang, Henan | 111.70 | 33.63 | 4 |
| LYG | 甘肃天水 Tianshui, Gansu | 106.10 | 34.23 | 5 |
| YPC | 甘肃陇南 Longnan, Gansu | 106.23 | 33.67 | 5 |
| HJG | 甘肃康县 Kang Xian, Gansu | 105.51 | 33.39 | 5 |
| MYG | 甘肃阳坝镇 Yangba, Gansu | 105.74 | 33.03 | 6 |
| SNJ | 湖北神农架 Shennongjia, Hubei | 110.92 | 31.65 | 7 |
| XJZ | 湖北神农架 Shennongjia, Hubei | 110.58 | 31.59 | 11 |
| SHJZ | 湖北恩施 Enshi, Hubei | 109.80 | 30.16 | 11 |
| JSZ | 重庆金山镇 Jinshan, Chongqing | 107.14 | 29.02 | 7 |
| DFX | 贵州毕节 Bijie, Guizhou | 105.88 | 27.33 | 12 |
| NYX | 贵州毕节 Bijie, Guizhou | 105.47 | 26.70 | 13 |

Fig. 2 Abnormal single nucleotide polymorphism (SNP) detected in Pterocarya hupehensis based on “Pcadapt” (A), Principal Component Analysis (PCA) (B), and latent factor mixed models analysis based on six environmental factors (C). The red dotted line represents the threshold of p = 0.01, and the sites above the red dotted line are significantly associated with the environmental factor. MAF, minor allele frequency.
| Mantel检验 Mantel test | Mantel’s r | p | Mantel检验(每个气候因子) Mantel test (Each climatic factor) | Mantel’s r | p |
|---|---|---|---|---|---|
| 地理隔离 Isolation by distance | 0.23 | 0.062 | 等温性 Isothermality | -0.17 | 0.861 |
| 环境隔离 Isolation by environment | 0.31 | 0.002 | 最冷月份最低气温 Minimum temperature of the coldest month | -0.05 | 0.668 |
| 偏Mantel检验 Partial mantel test | 气温年较差 Temperature annual range | -0.07 | 0.675 | ||
| 地理隔离(控制环境距离) Isolation by distance (Conditioned with environmental distance) | 0.08 | 0.298 | 最湿季度平均气温 Mean temperature of the wettest quarter | 0.09 | 0.212 |
| 环境隔离(控制地理距离) Isolation by environment (Conditioned with geographical distance) | 0.23 | 0.029 | 最湿月份降水量 Precipitation of the wettest month | 0.29 | 0.003 |
| 降水量季节性变化 Precipitation seasonality | 0.34 | 0.007 |
Table 2 Mantel and partial Mantel test results for the entire population of Pterocarya hupehensis
| Mantel检验 Mantel test | Mantel’s r | p | Mantel检验(每个气候因子) Mantel test (Each climatic factor) | Mantel’s r | p |
|---|---|---|---|---|---|
| 地理隔离 Isolation by distance | 0.23 | 0.062 | 等温性 Isothermality | -0.17 | 0.861 |
| 环境隔离 Isolation by environment | 0.31 | 0.002 | 最冷月份最低气温 Minimum temperature of the coldest month | -0.05 | 0.668 |
| 偏Mantel检验 Partial mantel test | 气温年较差 Temperature annual range | -0.07 | 0.675 | ||
| 地理隔离(控制环境距离) Isolation by distance (Conditioned with environmental distance) | 0.08 | 0.298 | 最湿季度平均气温 Mean temperature of the wettest quarter | 0.09 | 0.212 |
| 环境隔离(控制地理距离) Isolation by environment (Conditioned with geographical distance) | 0.23 | 0.029 | 最湿月份降水量 Precipitation of the wettest month | 0.29 | 0.003 |
| 降水量季节性变化 Precipitation seasonality | 0.34 | 0.007 |
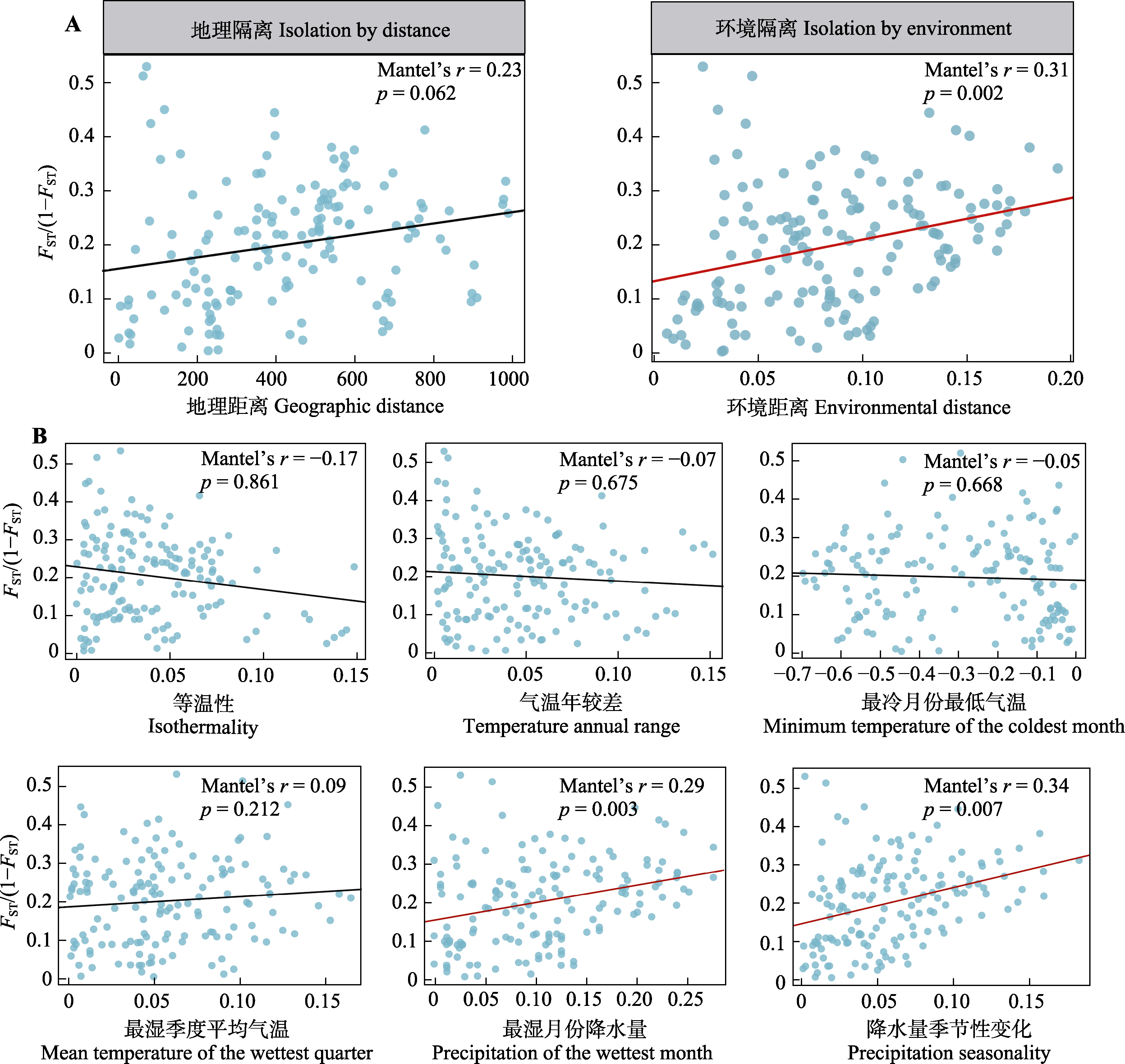
Fig. 3 Mantel test results showing the relationship between geographic distance and genetic distance, and between environmental distance and genetic distance across all populations (A), as well as the results of the genetic distance versus environmental distance for each individual climate factor (B). Red dashed lines represent significant correlations between variables. FST/(1 - FST), measuring genetic differentiation between paired populations (genetic distance).

Fig. 4 Redundancy analysis (RDA) of P. hupehensis between genetic variation and six climatic factors (A), and between genetic variation and geographical factors (B). As well as partial redundancy analysis (pRDA) between genetic variation and six climatic factors (C), and between genetic variation and geographical factors (D). The length of the vector represents the contribution of the environment variable to the explained variance, and the angle between the arrows represents the correlation between the variables. MEM, Moran’s eigenvector. bio3, isothermality; bio6, minimum temperature of the coldest month; bio7, temperature annual range; bio8, mean temperature of the wettest quarter; bio13, precipitation of the wettest month; bio15, precipitation seasonality.
| 环境因子 Environmental factor | 冗余分析 RDA | 偏冗余分析 Partial RDA | ||||
|---|---|---|---|---|---|---|
| PVE | 特征值 Eigenvalue | p | PVE | 特征值 Eigenvalue | p | |
| 地理 Geography | 0.13 | 3.54 | 0.001 | 0.09 | 2.54 | 0.001 |
| 气候 Climate | 0.18 | 4.06 | 0.001 | 0.13 | 3.17 | 0.001 |
| 等温性 Isothermality | 0.03 | 3.63 | 0.001 | 0.03 | 4.05 | 0.001 |
| 最冷月份最低气温 Minimum temperature of the coldest month | 0.03 | 3.70 | 0.001 | 0.02 | 2.60 | 0.001 |
| 气温年较差 Temperature annual range | 0.04 | 5.62 | 0.001 | 0.02 | 2.92 | 0.001 |
| 最湿季度平均气温 Mean temperature of the wettest quarter | 0.02 | 2.82 | 0.001 | 0.03 | 4.24 | 0.001 |
| 最湿月份降水量 Precipitation of the wettest month | 0.05 | 6.71 | 0.001 | 0.01 | 1.81 | 0.001 |
| 降水量季节性变化 Precipitation seasonality | 0.01 | 1.86 | 0.003 | 0.02 | 3.44 | 0.001 |
Table 3 Redundancy analysis (RDA) and partial RDA analysis results of all populations of Pterocarya hupehensis
| 环境因子 Environmental factor | 冗余分析 RDA | 偏冗余分析 Partial RDA | ||||
|---|---|---|---|---|---|---|
| PVE | 特征值 Eigenvalue | p | PVE | 特征值 Eigenvalue | p | |
| 地理 Geography | 0.13 | 3.54 | 0.001 | 0.09 | 2.54 | 0.001 |
| 气候 Climate | 0.18 | 4.06 | 0.001 | 0.13 | 3.17 | 0.001 |
| 等温性 Isothermality | 0.03 | 3.63 | 0.001 | 0.03 | 4.05 | 0.001 |
| 最冷月份最低气温 Minimum temperature of the coldest month | 0.03 | 3.70 | 0.001 | 0.02 | 2.60 | 0.001 |
| 气温年较差 Temperature annual range | 0.04 | 5.62 | 0.001 | 0.02 | 2.92 | 0.001 |
| 最湿季度平均气温 Mean temperature of the wettest quarter | 0.02 | 2.82 | 0.001 | 0.03 | 4.24 | 0.001 |
| 最湿月份降水量 Precipitation of the wettest month | 0.05 | 6.71 | 0.001 | 0.01 | 1.81 | 0.001 |
| 降水量季节性变化 Precipitation seasonality | 0.01 | 1.86 | 0.003 | 0.02 | 3.44 | 0.001 |
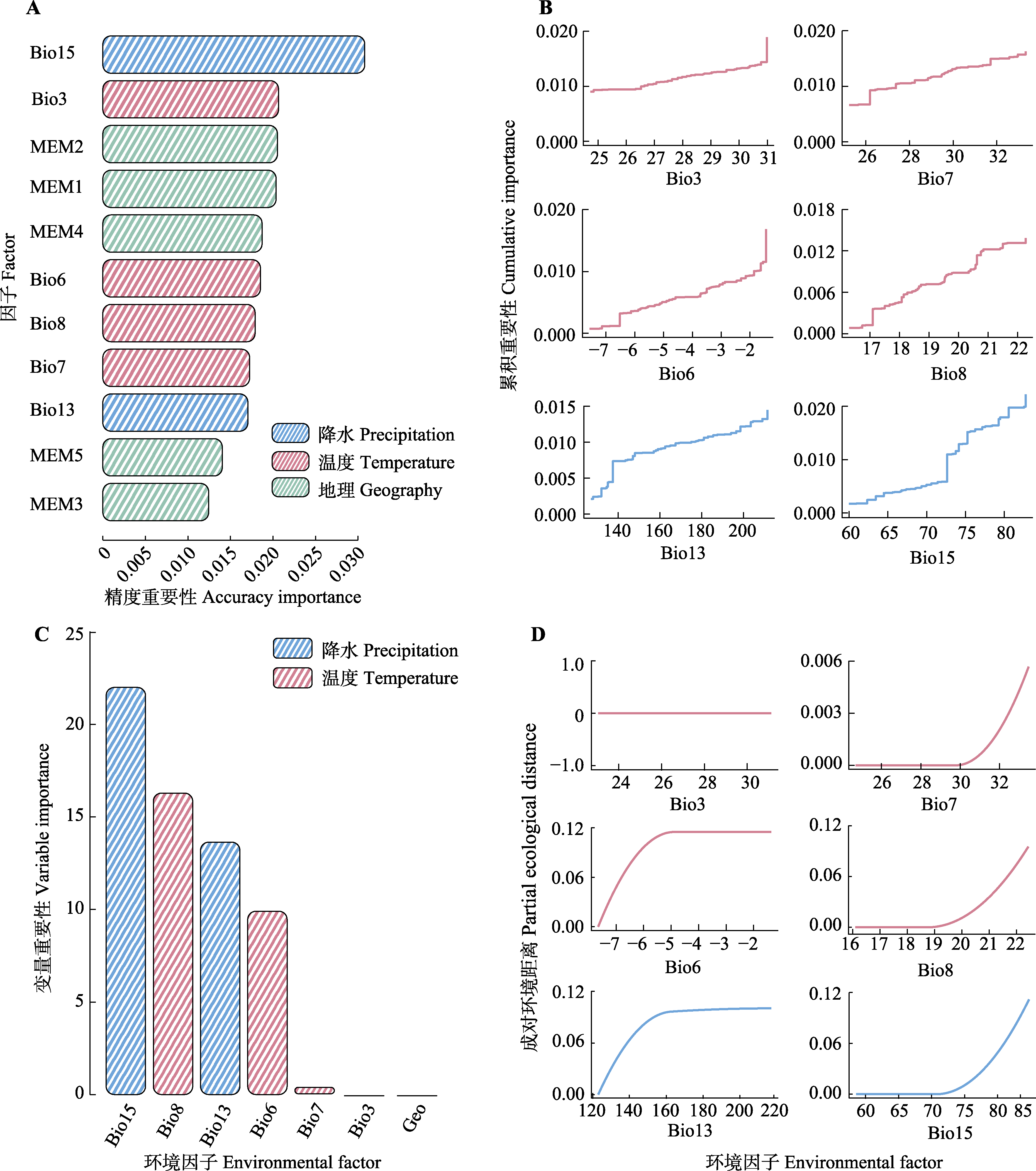
Fig. 5 Environmental variable importance rankings for genetic variation based on Gradient Forest (GF) (A) and Generalized Dissimilarity Model (GDM) (C) analyses, with I-spline curves illustrating genetic composition shifts along environmental gradients (B) and cumulative importance curves (D) in GDM analysis. MEM, Moran’s eigenvector. bio3, isothermality; bio6, minimum temperature of the coldest month; bio7, temperature annual range; bio8, mean temperature of the wettest quarter; bio13, precipitation of the wettest month; bio15, precipitation seasonality.
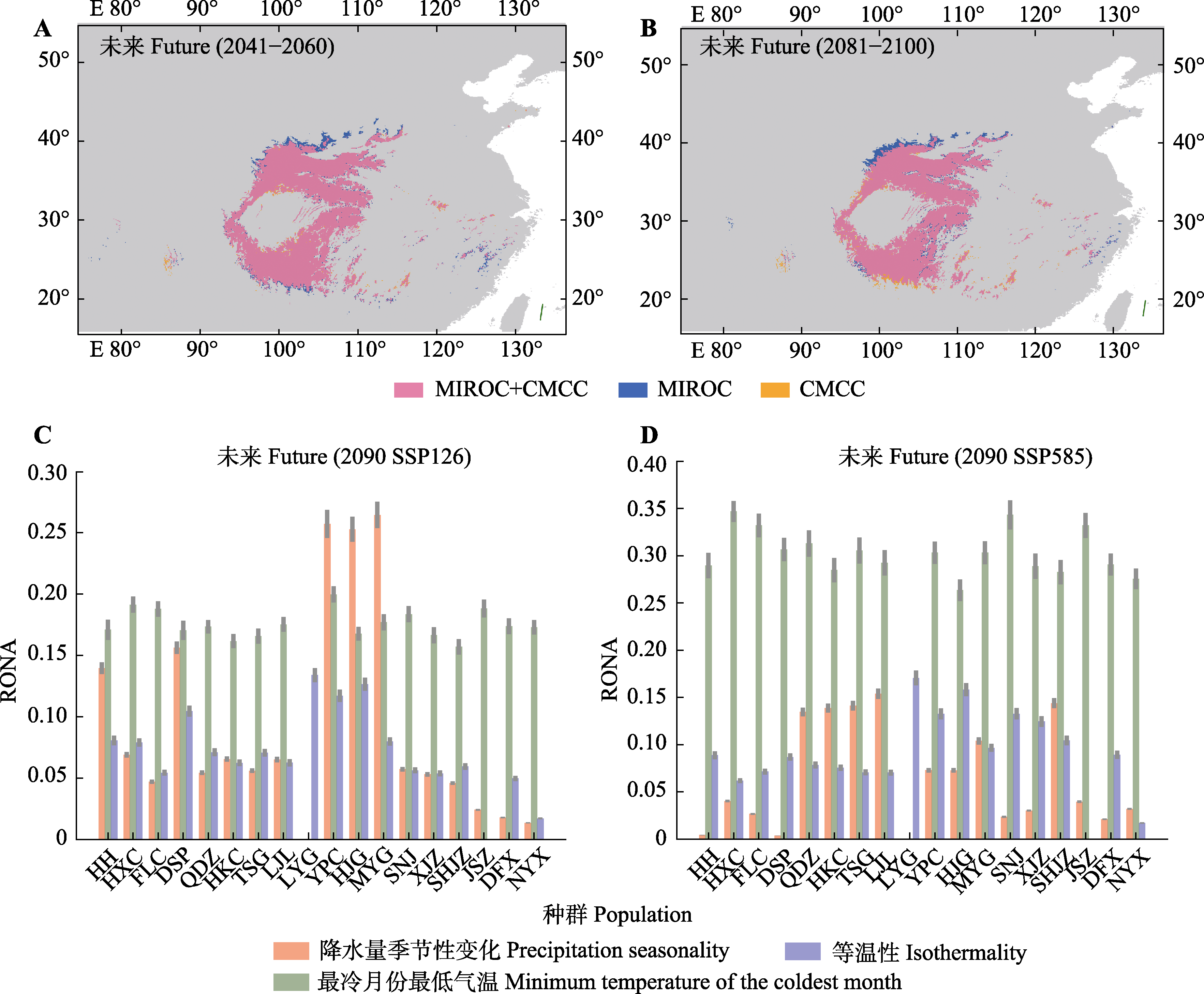
Fig. 6 Potential range projections (A, B) and risk of non-adaptedness analysis (RONA) (C, D) for future climate scenarios. Only the three climate factors that have the greatest impact on genomic vulnerability are shown in the picture. Data of two global climate models (MIROC and CMCC) were downloaded from WorldClim v2.1 (https://www.worldclim.org/data/cmip6/cmip6_clim2.5m. html). Detailed information of abbreviations for each population is shown in Table 1.
| [1] |
Aitken SN, Yeaman S, Holliday JA, Wang TL, Curtis-McLane S (2008). Adaptation, migration or extirpation: climate change outcomes for tree populations. Evolutionary Applications, 1, 95-111.
DOI PMID |
| [2] |
Anderson K, Gaston KJ (2013). Lightweight unmanned aerial vehicles will revolutionize spatial ecology. Frontiers in Ecology and the Environment, 11, 138-146.
DOI URL |
| [3] |
Andrews KR, Seaborn T, Egan JP, Fagnan MW, New DD, Chen ZQ, Hohenlohe PA, Waits LP, Caudill CC, Narum SR (2023). Whole genome resequencing identifies local adaptation associated with environmental variation for redband trout. Molecular Ecology, 32, 800-818.
DOI URL |
| [4] |
Bai WN, Wang WT, Zhang DY (2016). Phylogeographic breaks within Asian butternuts indicate the existence of a phytogeographic divide in East Asia. New Phytologist, 209, 1757-1772.
DOI URL |
| [5] | Balkenhol N, Cushman S, Storfer A, Waits L (2015). Landscape Genetics: Concepts, Methods, Applications. Wiley-Blackwell, Oxford, UK. |
| [6] | Balkenhol N, Dudaniec RY, Krutovsky KV, Johnson Jeremy S, Cairns DM, Segelbacher G, Selkoe KA, Heyden VDS, Wang IJ, Selmoni O, Joost Stéphane J (2019). Landscape genomics: understanding relationships between environmental heterogeneity and genomic characteristics of populations//Rajora OP. Population Genomics: Concepts, Approaches and Applications. Springer International Publishing, Cham, Switzerland. 261-322. |
| [7] |
Benito Garzón M, Robson TM, Hampe A (2019). ΔTraitSDMs: species distribution models that account for local adaptation and phenotypic plasticity. New Phytologist, 222, 1757-1765.
DOI PMID |
| [8] |
Cao YN, Comes HP, Sakaguchi S, Chen LY, Qiu YX (2016). Evolution of East Asia’s Arcto-Tertiary relict Euptelea (Eupteleaceae) shaped by Late Neogene vicariance and Quaternary climate change. BMC Evolutionary Biology, 16, 66.
DOI URL |
| [9] |
Capblancq T, Fitzpatrick MC, Bay RA, Exposito-Alonso M, Keller SR (2020). Genomic prediction of (mal)adaptation across current and future climatic landscapes. Annual Review of Ecology, Evolution, and Systematics, 51, 245-269.
DOI |
| [10] |
Catchen J, Hohenlohe PA, Bassham S, Amores A, Cresko WA (2013). Stacks: an analysis tool set for population genomics. Molecular Ecology, 22, 3124-3140.
DOI PMID |
| [11] |
Danecek P, Auton A, Abecasis G, Albers CA, Banks E, DePristo MA, Handsaker RE, Lunter G, Marth GT, Sherry ST, McVean G, Durbin R,1000 Genomes Project Analysis Group (2011). The variant call format and VCFtools. Bioinformatics, 27, 2156-2158.
DOI PMID |
| [12] |
Dauphin B, Rellstab C, Schmid M, Zoller S, Karger DN, Brodbeck S, Guillaume F, Gugerli F (2021). Genomic vulnerability to rapid climate warming in a tree species with a long generation time. Global Change Biology, 27, 1181-1195.
DOI PMID |
| [13] |
Diniz-Filho JAF, Soares TN, Lima JS, Dobrovolski R, Landeiro VL, de Campos Telles MP, Rangel TF, Bini LM (2013). Mantel test in population genetics. Genetics and Molecular Biology, 36, 475-485.
DOI PMID |
| [14] | Doyle JJ, Doyle JL (1987). A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochemical Bulletin, 19, 11-15. |
| [15] |
Ellis N, Smith SJ, Pitcher CR (2012). Gradient forests: calculating importance gradients on physical predictors. Ecology, 93, 156-168.
PMID |
| [16] | Fahad S, Sonmez O, Saud S, Wang D, Wu C, Adnan M, Turan V (2021). Climate Change and Plants: Biodiversity, Growth and Interactions. CRC Press, Boca Raton, USA. |
| [17] |
Ferrier S, Manion G, Elith J, Richardson K (2007). Using generalized dissimilarity modelling to analyse and predict patterns of beta diversity in regional biodiversity assessment. Diversity and Distributions, 13, 252-264.
DOI URL |
| [18] |
Fick SE, Hijmans RJ (2017). WorldClim 2: new 1-km spatial resolution climate surfaces for global land areas. International Journal of Climatology, 37, 4302-4315.
DOI URL |
| [19] | Foden WB, Young BE, Akçakaya HR, Garcia RA, Hoffmann AA, Stein BA, Thomas CD, Wheatley CJ, Bickford D, Carr JA, Hole DG, Martin TG, Pacifici M, Pearce-Higgins JW, Platts PJ, et al. (2019). Climate change vulnerability assessment of species. Wiley Interdisciplinary Reviews: Climate Change, 10, e551. DOI: 10.1002/wcc.551. |
| [20] | Frankel OH, Brown AHD, Burdon J (1995). The genetic diversity of wild plants//Volin VC, Volin J. The Conservation of Plant Biodiversity. Cambridge University Press, Cambridge, UK. 10-38. |
| [21] |
Frichot E, François O (2015). LEA: an R package for landscape and ecological association studies. Methods in Ecology and Evolution, 6, 925-929.
DOI URL |
| [22] | Goslee SC, Urban DL (2007). The ecodist package for dissimilarity-based analysis of ecological data. Journal of Statistical Software, 22, 1-19. |
| [23] |
Gougherty AV, Keller SR, Fitzpatrick MC (2021). Maladaptation, migration and extirpation fuel climate change risk in a forest tree species. Nature Climate Change, 11, 166-171.
DOI |
| [24] |
Gugger PF, Liang CT, Sork VL, Hodgskiss P, Wright JW (2018). Applying landscape genomic tools to forest management and restoration of Hawaiian koa (Acacia koa) in a changing environment. Evolutionary Applications, 11, 231-242.
DOI PMID |
| [25] |
Hijmans RJ, Cameron SE, Parra JL, Jones PG, Jarvis A (2005). Very high resolution interpolated climate surfaces for global land areas. International Journal of Climatology, 25, 1965-1978.
DOI URL |
| [26] | Hijmans RJ, Phillips S, Leathwick J, Elith J (2017a). dismo: Species distribution modeling. R package version 1. 1-4. [2024-07-02]. https://CRAN.R-project.org/package=dismo. |
| [27] | Hijmans RJ, van Etten J, Cheng J, Mattiuzzi M, Sumner M, Greenberg JA, Lamigueiro OP, Bevan A, Racine EB, Shortridge A, Ghosh A (2017b). Package ‘raster’: geographic data analysis and modeling, v. 2.6-7. [2024-07-08]. https://cran.r-project.org/web/packages/raster/raster.pdf. |
| [28] | Hijmans RJ, Williams E, Vennes C, Hijmans MR (2021). Package ‘geosphere’. R package version 1.5-14. [2024-06-01]. https://cran.r-project.org/web/packages/geosphere/index.html. |
| [29] |
Hitchings SP, Beebee TJC (1997). Genetic substructuring as a result of barriers to gene flow in urban Rana temporaria (common frog) populations: implications for biodiversity conservation. Heredity, 79, 117-127.
DOI |
| [30] |
Holliday JA, Aitken SN, Cooke JEK, Fady B, González-Martínez SC, Heuertz M, Jaramillo-Correa JP, Lexer C, Staton M, Whetten RW, Plomion C (2017). Advances in ecological genomics in forest trees and applications to genetic resources conservation and breeding. Molecular Ecology, 26, 706-717.
DOI PMID |
| [31] | Hurtt GC, Chini L, Sahajpal R, Frolking S, Bodirsky BL, Calvin K, Doelman JC, Fisk J, Fujimori S, Klein Goldewijk K, Hasegawa T, Havlik P, Heinimann A, Humpenöder F, Jungclaus J, et al. (2020). Harmonization of global land use change and management for the period 850-2100 (LUH2) for CMIP6. Geoscientific Model Development, 13, 5425-5464. |
| [32] | Jiang S, Luo MX, Gao RH, Zhang W, Yang YZ, Li YJ, Liao PC (2019). Isolation-by-environment as a driver of genetic differentiation among populations of the only broad-leaved evergreen shrub Ammopiptanthus mongolicus in Asian temperate deserts. Scientific Reports, 9, 12008. DOI: 10.1038/s41598-019-48472-y. |
| [33] |
Kou YX, Cheng SM, Tian S, Li B, Fan DM, Chen YJ, Soltis DE, Soltis PS, Zhang ZY (2016). The antiquity of Cyclocarya paliurus (Juglandaceae) provides new insights into the evolution of relict plants in subtropical China since the late Early Miocene. Journal of Biogeography, 43, 351-360.
DOI URL |
| [34] | Kozlowski G, Sébastien B, Song YG (2018). Wingnuts (Pterocarya) & Walnut Family. Relict Trees: Linking the Past, Present and Future. Natural History Museum Fribourg, Fribourg, Switzerland. |
| [35] | Kozlowski G, Song Y, Bétrisey S (2019). Pterocarya hupehensis. The IUCN Red List of Threatened Species 2019: e. T66816108A152835141. [2024-09-12]. http://dx. doi.org/10.2305/IUCN.UK.2019-3.RLTS.T66816108A152835141.en. |
| [36] | Kuang KR, Li PQ (1979). Flora of China: Volume 22. Science Press, Beijing. |
| [匡可任, 李沛琼 (1979). 中国植物志: 第二十二卷. 科学出版社, 北京.] | |
| [37] |
Lasky JR, des Marais DL, McKay JK, Richards JH, Juenger TE, Keitt TH (2012). Characterizing genomic variation of Arabidopsis thaliana: the roles of geography and climate. Molecular Ecology, 21, 5512-5529.
DOI URL |
| [38] |
Lefèvre F, Boivin T, Bontemps A, Courbet F, Davi H, Durand-Gillmann M, Fady B, Gauzere J, Gidoin C, Karam MJ, Lalagüe H, Oddou-Muratorio S, Pichot C (2014). Considering evolutionary processes in adaptive forestry. Annals of Forest Science, 71, 723-739.
DOI URL |
| [39] |
Legendre P, Anderson MJ (1999). Distance-based redundancy analysis: testing multispecies responses in multifactorial ecological experiments. Ecological Monographs, 69, 1-24.
DOI URL |
| [40] |
Li LF, Cushman SA, He YX, Ma XF, Ge XJ, Li JX, Qian ZH, Li Y (2022). Landscape genomics reveals genetic evidence of local adaptation in a widespread tree, the Chinese wingnut (Pterocarya stenoptera). Journal of Systematics and Evolution, 60, 386-397.
DOI URL |
| [41] | Liaw A, Wiener M (2002). Classification and Regression by randomForest. R News, 2, 18-22. |
| [42] |
Lischer HEL, Excoffier L (2012). PGDSpider: an automated data conversion tool for connecting population genetics and genomics programs. Bioinformatics, 28, 298-299.
DOI PMID |
| [43] | Lu ZJ, Wang TR, Zheng SS, Meng HH, Cao JG, Song YG, Kozlowski G (2024). Phylogeography of Pterocarya hupehensis reveals the evolutionary patterns of a Cenozoic relict tree around the Sichuan Basin. Forestry Research, 4, e008. DOI: 10.48130/forres-0024-0005. |
| [44] |
Luu K, Bazin E, Blum MGB (2017). Pcadapt: an R package to perform genome scans for selection based on principal component analysis. Molecular Ecology Resources, 17, 67-77.
DOI PMID |
| [45] |
Manel S, Schwartz MK, Luikart G, Taberlet P (2003). Landscape genetics: combining landscape ecology and population genetics. Trends in Ecology & Evolution, 18, 189-197.
DOI URL |
| [46] |
Meinshausen M, Nicholls ZRJ, Lewis J, Gidden MJ, Vogel E, Freund M, Beyerle U, Gessner C, Nauels A, Bauer N, Canadell JG, Daniel JS, John A, Krummel PB, Luderer G, et al. (2020). The shared socio-economic pathway (SSP) greenhouse gas concentrations and their extensions to 2500. Geoscientific Model Development, 13, 3571-3605.
DOI |
| [47] |
Meng HH, Gao XY, Song YG, Cao GL, Li J (2021). Biodiversity arks in the anthropocene. Regional Sustainability, 2, 109-115.
DOI |
| [48] |
Naimi B, Hamm NAS, Groen TA, Skidmore AK, Toxopeus AG (2014). Where is positional uncertainty a problem for species distribution modelling? Ecography, 37, 191-203.
DOI URL |
| [49] |
Pacifici M, Foden WB, Visconti P, Watson JEM, Butchart SHM, Kovacs KM, Scheffers BR, Hole DG, Martin TG, Akçakaya HR, Corlett RT, Huntley B, Bickford D, Carr JA, Hoffmann AA, et al. (2015). Assessing species vulnerability to climate change. Nature Climate Change, 5, 215-224.
DOI |
| [50] |
Pettorelli N, Vik JO, Mysterud A, Gaillard JM, Tucker CJ, Stenseth NC (2005). Using the satellite-derived NDVI to assess ecological responses to environmental change. Trends in Ecology & Evolution, 20, 503-510.
DOI URL |
| [51] |
Phillips SJ, Anderson RP, Schapire RE (2006). Maximum entropy modeling of species geographic distributions. Ecological Modelling, 190, 231-259.
DOI URL |
| [52] |
Pina-Martins F, Baptista J, Pappas G, Paulo OS (2019). New insights into adaptation and population structure of cork oak using genotyping by sequencing. Global Change Biology, 25, 337-350.
DOI PMID |
| [53] |
Privé F, Luu K, Vilhjálmsson BJ, Blum MGB (2020). Performing highly efficient genome scans for local adaptation with R package pcadapt version 4. Molecular Biology and Evolution, 37, 2153-2154.
DOI PMID |
| [54] |
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MAR, Bender D, Maller J, Sklar P, Daly MJ, Sham PC (2007). PLINK: a tool set for whole-genome association and population-based linkage analyses. American Journal of Human Genetics, 81, 559-575.
DOI PMID |
| [55] |
Raymond M, Rousset F (1995). GENEPOP (version 1.2): population genetics software for exact tests and ecumenicism. Journal of Heredity, 86, 248-249.
DOI URL |
| [56] |
Rellstab C, Gugerli F, Eckert AJ, Hancock AM, Holderegger R (2015). A practical guide to environmental association analysis in landscape genomics. Molecular Ecology, 24, 4348-4370.
DOI PMID |
| [57] |
Rellstab C, Zoller S, Walthert L, Lesur I, Pluess AR, Graf R, Bodénès C, Sperisen C, Kremer A, Gugerli F (2016). Signatures of local adaptation in candidate genes of oaks (Quercus spp.) with respect to present and future climatic conditions. Molecular Ecology, 25, 5907-5924.
DOI URL |
| [58] |
Root TL, Price JT, Hall KR, Schneider SH, Rosenzweig C,Alan Pounds J (2003). Fingerprints of global warming on wild animals and plants. Nature, 421, 57-60.
DOI |
| [59] |
Rousset F (2008). Genepop’007: a complete re-implementation of the genepop software for Windows and Linux. Molecular Ecology Resources, 8, 103-106.
DOI URL |
| [60] | Sang YP, Long ZQ, Dan XM, Feng JJ, Shi TT, Jia CF, Zhang XX, Lai Q, Yang GL, Zhang HY, Xu XT, Liu HH, Jiang YZ, Ingvarsson PK, Liu JQ, Mao KS, Wang J (2022). Genomic insights into local adaptation and future climate-induced vulnerability of a keystone forest tree in East Asia. Nature Communications, 13, 6541. DOI: 10.1038/s41467-022-34206-8. |
| [61] |
Savolainen O, Pyhäjärvi T, Knürr T (2007). Gene flow and local adaptation in trees. Annual Review of Ecology, Evolution, and Systematics, 38, 595-619.
DOI URL |
| [62] | Scheffers BR, De Meester L, Bridge TCL, Hoffmann AA, Pandolfi JM, Corlett RT, Butchart SHM, Pearce-Kelly P, Kovacs KM, Dudgeon D, Pacifici M, Rondinini C, Foden WB, Martin TG, Mora C, et al. (2016). The broad footprint of climate change from genes to biomes to people. Science, 354, aaf7671. DOI: 10.1126/science.aaf7671. |
| [63] | Smith SJ, Ellis N, Pitcher CR (2011). Conditional variable importance in R package extendedForest. [2024-06-06]. https://gradientforest.r-forge.r-project.org/Conditional-importance.pdf. |
| [64] |
Sork VL, Waits L (2010). Contributions of landscape genetics-approaches, insights, and future potential. Molecular Ecology, 19, 3489-3495.
DOI URL |
| [65] |
Storfer A, Murphy MA, Evans JS, Goldberg CS, Robinson S, Spear SF, Dezzani R, Delmelle E, Vierling L, Waits LP (2007). Putting the ‘landscape’ in landscape genetics. Heredity, 98, 128-142.
PMID |
| [66] |
Thuiller W, Albert C, Araújo MB, Berry PM, Cabeza M, Guisan A, Hickler T, Midgley GF, Paterson J, Schurr FM, Sykes MT, Zimmermann NE (2008). Predicting global change impacts on plant species’ distributions: future challenges. Perspectives in Plant Ecology, Evolution and Systematics, 9, 137-152.
DOI URL |
| [67] |
Urban MC (2015). Accelerating extinction risk from climate change. Science, 348, 571-573.
DOI PMID |
| [68] |
van Strien MJ, Holderegger R, Van Heck HJ (2015). Isolation-by-distance in landscapes: considerations for landscape genetics. Heredity, 114, 27-37.
DOI PMID |
| [69] |
Wahid A, Gelani S, Ashraf M, Foolad MR (2007). Heat tolerance in plants: an overview. Environmental and Experimental Botany, 61, 199-223.
DOI URL |
| [70] |
Wang IJ, Bradburd GS (2014). Isolation by environment. Molecular Ecology, 23, 5649-5662.
DOI PMID |
| [71] | Wang TR, Feng L, Du F (2021). New approaches for ecological adaptation study: from population genetics to landscape genomics. Scientia Sinica (Vitae), 51, 167-178. |
| [王天瑞, 冯力, 杜芳 (2021). 生态适应研究新方法: 从种群遗传学到景观基因组学. 中国科学: 生命科学, 51, 167-178. | |
| [72] | Wang TR, Meng HH, Wang N, Zheng SS, Jiang Y, Lin DQ, Song YG, Kozlowski G (2023). Adaptive divergence and genetic vulnerability of relict species under climate change: a case study of Pterocarya macroptera. Annals of Botany, 132, 241-254. |
| [73] | Wang TR, Qiu YX (2024). Application and prospects of landscape genomics in conservation biology. Plant Science Journal, 41, 741-750. |
| [王天瑞, 邱英雄 (2024). 景观基因组学方法在保护生物学研究中的应用与前景. 植物科学学报, 41, 741-750.] | |
| [74] |
Yin QY, Fan Q, Li P, Truong D, Zhao WY, Zhou RC, Chen SF, Liao WB (2021). Neogene and Quaternary climate changes shaped the lineage differentiation and demographic history of Fokienia hodginsii (Cupressaceae s.l.), a Tertiary relict in East Asia. Journal of Systematics and Evolution, 59, 1081-1099.
DOI URL |
| [75] | Zhang WP, Cao L, Lin XR, Ding YM, Liang Y, Zhang DY, Pang EL, Renner SS, Bai WN (2022). Dead-end hybridization in walnut trees revealed by large-scale genomic sequence data. Molecular Biology and Evolution, 39, msab308. DOI: 10.1093/molbev/msab308. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2026 Chinese Journal of Plant Ecology
Tel: 010-62836134, 62836138, E-mail: apes@ibcas.ac.cn, cjpe@ibcas.ac.cn
![]()