

植物生态学报 ›› 2025, Vol. 49 ›› Issue (11): 1858-1868.DOI: 10.17521/cjpe.2024.0214 cstr: 32100.14.cjpe.2024.0214
收稿日期:2024-07-02
接受日期:2025-01-09
出版日期:2025-11-20
发布日期:2025-11-20
通讯作者:
*娄安如(louanru@bun.edu.cn)基金资助:
ZHANG Wang, TAN Si-Yi, TU Wen-Qin, LOU An-Ru*( )
)
Received:2024-07-02
Accepted:2025-01-09
Online:2025-11-20
Published:2025-11-20
Supported by:摘要:
该研究以新疆吐鲁番和哈密两个地区(以下简称吐哈地区)的9个梭梭(Haloxylon ammodendron)种群为研究对象, 结合准噶尔盆地东南缘昌吉州奇台县和木垒县2个种群, 以及甘肃省2个种群, 使用基因组重测序获得的单核苷酸多态性(SNP)数据和叶绿体基因组数据, 分析其遗传多样性及遗传结构。SNP数据分析表明, 新疆昌吉州奇台县的梭梭种群核苷酸多样性较高, 而其东部哈密市巴里坤哈萨克自治县以及伊州区北部西山乡、沁城乡等地的梭梭种群核苷酸多样性相对较低; 吐哈地区的9个梭梭种群可分为4个类群, 其中哈密市巴里坤县和镜儿泉的梭梭种群与准噶尔盆地东南缘种群和甘肃种群具有相似的遗传成分; 吐鲁番高昌区、哈密市西山乡分布的2个类群保留有独特的遗传成分。13个种群共有39个叶绿体单倍型, 其中新疆昌吉州奇台县单倍型最多且最古老, 多样性最高; 吐哈地区和甘肃种群中均含有奇台单倍型。综合核基因组和叶绿体基因组种群遗传格局研究结果, 支持研究区域梭梭种群由准噶尔盆地东南缘的奇台种群向东传播的趋势判断, 提出了吐哈地区梭梭保护建议。
张望, 谭思仪, 涂文琴, 娄安如. 新疆吐鲁番和哈密地区梭梭种群遗传多样性和遗传结构. 植物生态学报, 2025, 49(11): 1858-1868. DOI: 10.17521/cjpe.2024.0214
ZHANG Wang, TAN Si-Yi, TU Wen-Qin, LOU An-Ru. Genetic diversity and genetic structure of Haloxylon ammodendron in the Turpan and Hami area, Xinjiang, China. Chinese Journal of Plant Ecology, 2025, 49(11): 1858-1868. DOI: 10.17521/cjpe.2024.0214
| 序号 ID | 地点 Locality | 编号 Code | 经度 Longitude (E) | 纬度 Latitude (N) | 样本量 Sample size | 单倍型 Haplotypes | 核苷酸多样性 Nucleotide diversity (π) | Tajima D | 单倍型多样性 Haplotype diversity (Hd) | ||
|---|---|---|---|---|---|---|---|---|---|---|---|
| 新疆昌吉回族自治州 Changji Hui Autonomous Prefecture, Xinjiang | |||||||||||
| 1 | 奇台 Qitai | QT | 89.63° | 44.19° | 12 | H3(3), H4(1), H5(2), H6(1), H14(1), H15(1), H34(1), H35(1), 36(1) | 0.004 | -0.604 | 0.929 | ||
| 2 | 木垒 Mori | ML | 90.78° | 44.03° | 12 | H11(10), H29(1), H30(1) | 0.003 | -0.050 | 0.318 | ||
| 新疆吐鲁番和哈密地区 Turpan and Hami area, Xinjiang | |||||||||||
| 3 | 星星峡 Xingxingxia | XXX | 95.17° | 41.72° | 12 | H20(10) | 0.003 | 0.440 | 0.000 | ||
| 4 | 镜儿泉 Jing’erquan | JEQ | 95.34° | 42.35° | 12 | H7(1), H8(9), H28(1) | 0.004 | 0.194 | 0.345 | ||
| 5 | 沁城 Qincheng | QCN | 94.61° | 42.60° | 12 | H6(3), H12(2), H13(2), H14(1), H31(1), H32(1), H33(1) | 0.003 | -0.748 | 0.909 | ||
| 6 | 西山乡 Xishan Township | XSN | 93.35° | 43.06° | 12 | H5(1), H18(8), H19(1), H20(1) | 0.003 | 0.419 | 0.491 | ||
| 7 | 亚勒曼 Yaleman | YLM | 95.70° | 43.05° | 12 | H8(7), H40(5) | 0.003 | -0.010 | 0.530 | ||
| 8 | 三塘湖 Santanghu | STH | 93.21° | 44.11° | 12 | H6(7), H15(1), H16(1), H37(1), H38(1), 39(1) | 0.003 | -0.771 | 0.682 | ||
| 9 | 老爷庙 Laoyemiao | LYM | 93.93° | 44.66° | 12 | H9(7), H10(5) | 0.003 | -0.164 | 0.530 | ||
| 10 | 汉水泉 Hanshuiquan | HSQ | 92.05° | 44.52° | 12 | H6(4), H24(4), H25(1), H26(1), H27(1) | 0.002 | -0.843 | 0.782 | ||
| 11 | 吐鲁番 Turpan | TLF | 88.86° | 43.28° | 12 | H17(11), H18(1) | 0.004 | 0.493 | 0.167 | ||
| 甘肃 Gansu | |||||||||||
| 12 | 敦煌 Dunhuang | DH | 94.70° | 40.11° | 12 | H3(3), H4(4), H5(4) | 0.004 | -0.292 | 0.727 | ||
| 13 | 阿克塞 Aksay | AKS | 94.36° | 39.72° | 12 | H1(8), H2(2), H22(1), H23(1) | 0.003 | -0.630 | 0.561 | ||
表1 本研究中13个梭梭种群采样位置和遗传多样性信息
Table 1 Sampling information and genetic diversity of the 13 Haloxylon ammodendron populations
| 序号 ID | 地点 Locality | 编号 Code | 经度 Longitude (E) | 纬度 Latitude (N) | 样本量 Sample size | 单倍型 Haplotypes | 核苷酸多样性 Nucleotide diversity (π) | Tajima D | 单倍型多样性 Haplotype diversity (Hd) | ||
|---|---|---|---|---|---|---|---|---|---|---|---|
| 新疆昌吉回族自治州 Changji Hui Autonomous Prefecture, Xinjiang | |||||||||||
| 1 | 奇台 Qitai | QT | 89.63° | 44.19° | 12 | H3(3), H4(1), H5(2), H6(1), H14(1), H15(1), H34(1), H35(1), 36(1) | 0.004 | -0.604 | 0.929 | ||
| 2 | 木垒 Mori | ML | 90.78° | 44.03° | 12 | H11(10), H29(1), H30(1) | 0.003 | -0.050 | 0.318 | ||
| 新疆吐鲁番和哈密地区 Turpan and Hami area, Xinjiang | |||||||||||
| 3 | 星星峡 Xingxingxia | XXX | 95.17° | 41.72° | 12 | H20(10) | 0.003 | 0.440 | 0.000 | ||
| 4 | 镜儿泉 Jing’erquan | JEQ | 95.34° | 42.35° | 12 | H7(1), H8(9), H28(1) | 0.004 | 0.194 | 0.345 | ||
| 5 | 沁城 Qincheng | QCN | 94.61° | 42.60° | 12 | H6(3), H12(2), H13(2), H14(1), H31(1), H32(1), H33(1) | 0.003 | -0.748 | 0.909 | ||
| 6 | 西山乡 Xishan Township | XSN | 93.35° | 43.06° | 12 | H5(1), H18(8), H19(1), H20(1) | 0.003 | 0.419 | 0.491 | ||
| 7 | 亚勒曼 Yaleman | YLM | 95.70° | 43.05° | 12 | H8(7), H40(5) | 0.003 | -0.010 | 0.530 | ||
| 8 | 三塘湖 Santanghu | STH | 93.21° | 44.11° | 12 | H6(7), H15(1), H16(1), H37(1), H38(1), 39(1) | 0.003 | -0.771 | 0.682 | ||
| 9 | 老爷庙 Laoyemiao | LYM | 93.93° | 44.66° | 12 | H9(7), H10(5) | 0.003 | -0.164 | 0.530 | ||
| 10 | 汉水泉 Hanshuiquan | HSQ | 92.05° | 44.52° | 12 | H6(4), H24(4), H25(1), H26(1), H27(1) | 0.002 | -0.843 | 0.782 | ||
| 11 | 吐鲁番 Turpan | TLF | 88.86° | 43.28° | 12 | H17(11), H18(1) | 0.004 | 0.493 | 0.167 | ||
| 甘肃 Gansu | |||||||||||
| 12 | 敦煌 Dunhuang | DH | 94.70° | 40.11° | 12 | H3(3), H4(4), H5(4) | 0.004 | -0.292 | 0.727 | ||
| 13 | 阿克塞 Aksay | AKS | 94.36° | 39.72° | 12 | H1(8), H2(2), H22(1), H23(1) | 0.003 | -0.630 | 0.561 | ||
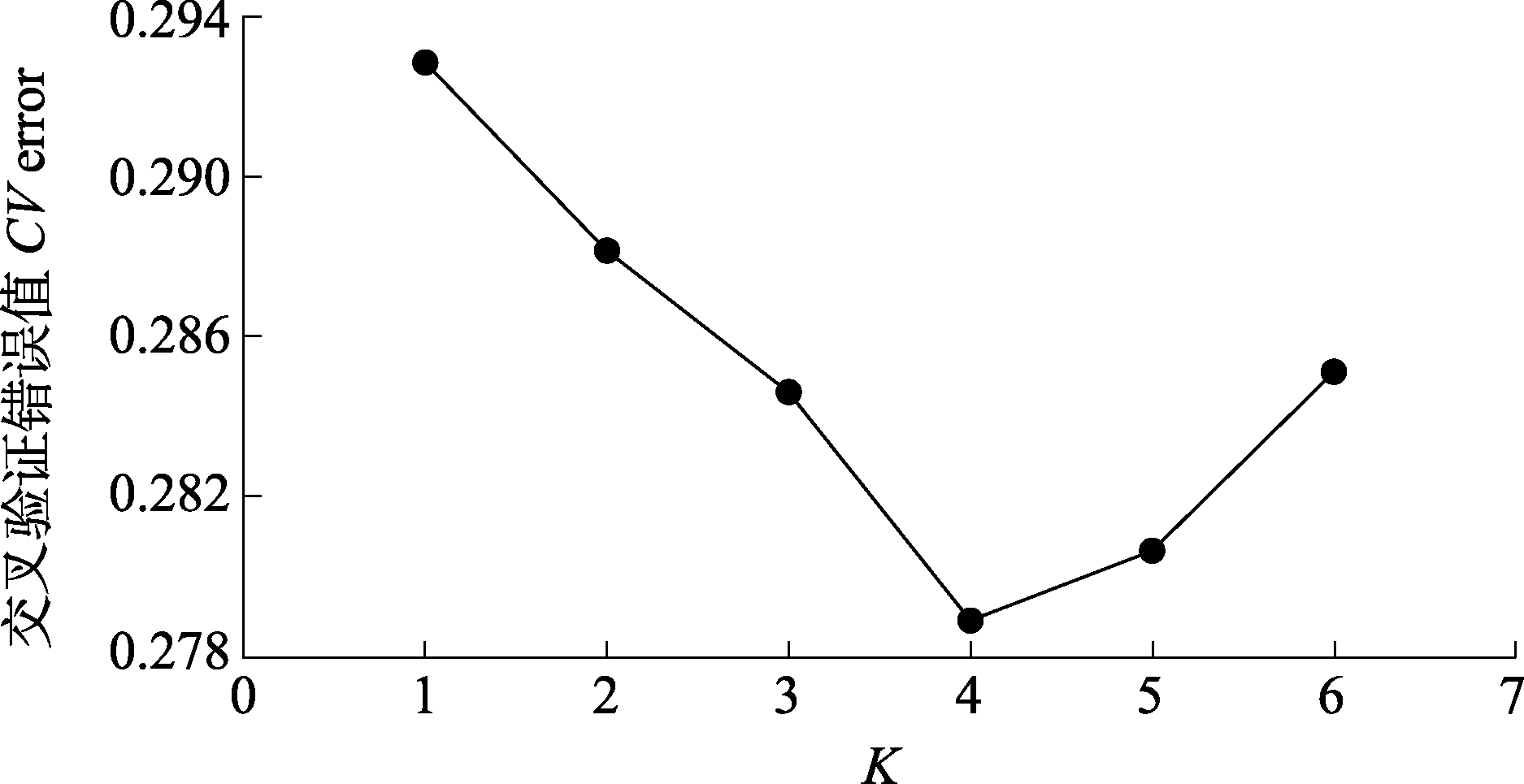
图1 13个梭梭种群K值为1-6时交叉验证错误值(CV error)。
Fig. 1 Estimation of cross-validation errors (CV error) for K values ranging from 1 to 6 of the13 Haloxylon ammodendron populations.
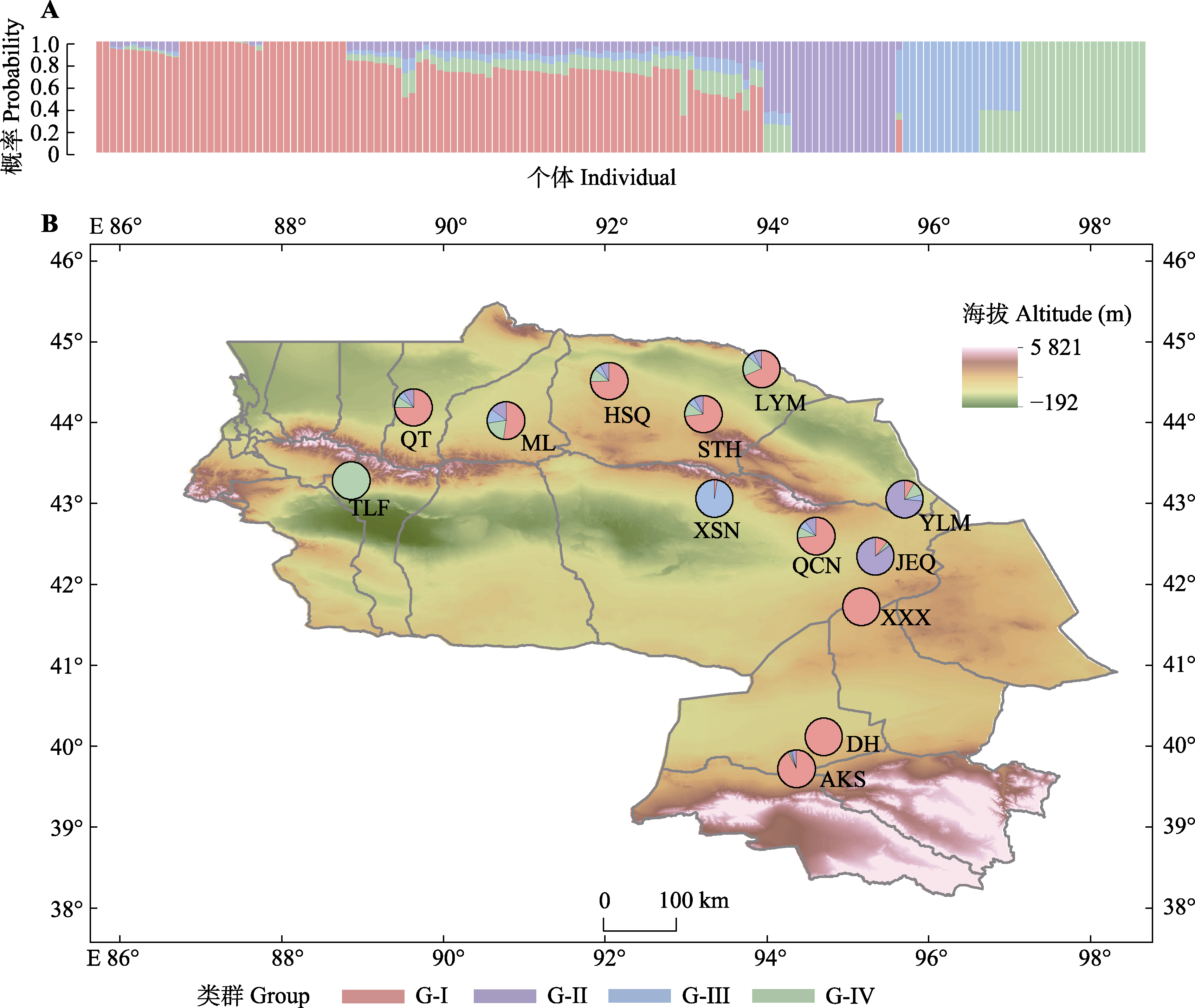
图2 基于单核苷酸多态性(SNP)分子标记的梭梭种群的遗传结构和地理分布。A, 13个梭梭种群的151个样本在K = 4时的ADMIXTURE结果图。B, 基于K = 4的13个梭梭种群的遗传结构地理分布图, 图中缩写为种群编号, 同表1。
Fig. 2 Results of population genetic structure and geographical distribution of Haloxylon ammodendron obtained based on single nucleotide polymerphism (SNP) markers. A, ADMIXTURE results for 151 samples from 13 populations at K = 4. B, Geographical distribution of the genetic structure of 13 H. ammodendron populations with ADMIXTURE at K = 4. The abbreviations indicate population codes, as shown in Table 1.
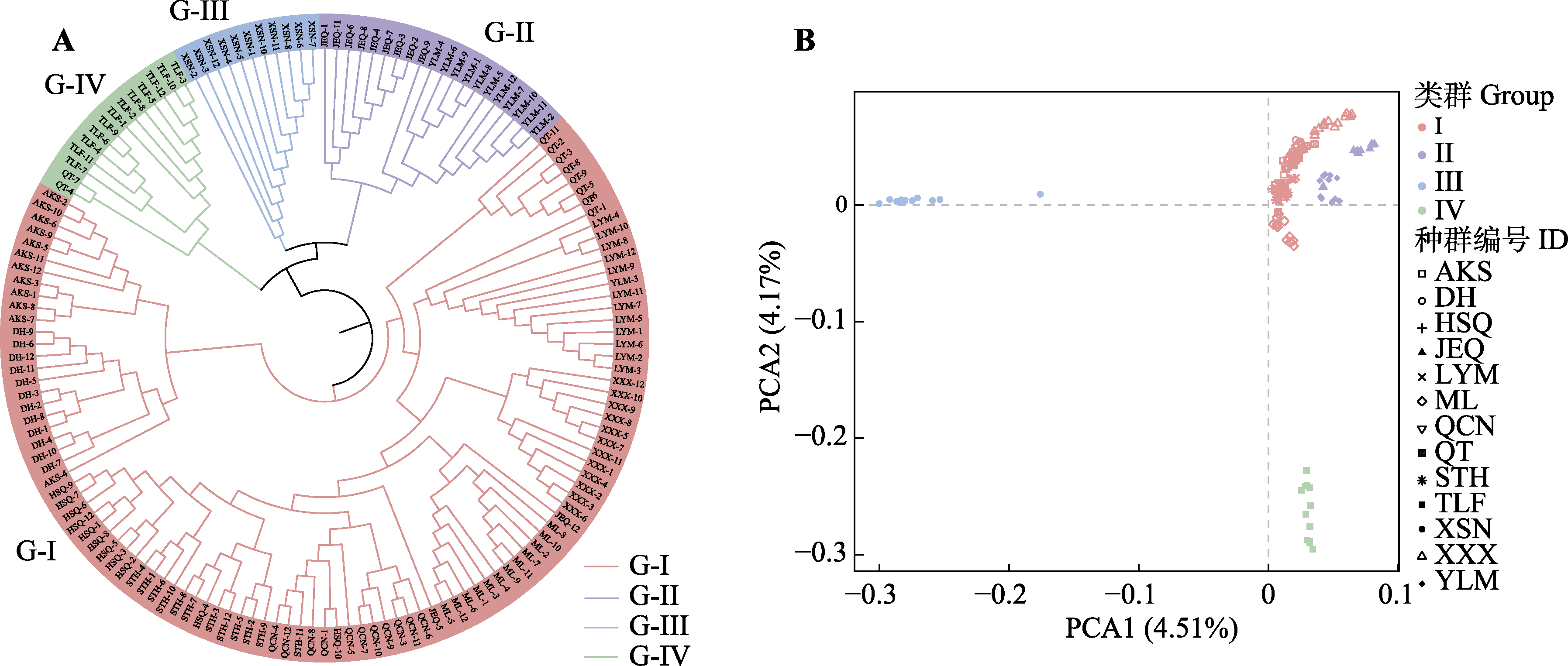
图3 基于单核苷酸多态性分子标记使用邻接法构建13个梭梭种群聚类树(A)和主成分分析(PCA)结果(B)。G-I-G-IV表示按照ADMIXTURE结果划分的四个类群。种群编号同表1。
Fig. 3 Neighbor-Joining tree (A) and principal components analysis (PCA) (B) of 13 Haloxylon ammodendron populations obtained based on single nucleotide polymorphism markers. G-I-G-IV represents the four groups divided according to the ADMIXTURE result. The abbreviations indicate population codes, as shown in Table 1.
| 类群 Inferred group | G-I | G-II | G-III | G-IV |
|---|---|---|---|---|
| G-I | - | 0.048 | 0.110 | 0.038 |
| G-II | - | 0.136 | 0.059 | |
| G-III | - | 0.118 | ||
| G-IV | - |
表2 4个梭梭类群间遗传分化(Fst)
Table 2 Pairwise genetic differentiation (Fst) of the four groups in this study
| 类群 Inferred group | G-I | G-II | G-III | G-IV |
|---|---|---|---|---|
| G-I | - | 0.048 | 0.110 | 0.038 |
| G-II | - | 0.136 | 0.059 | |
| G-III | - | 0.118 | ||
| G-IV | - |
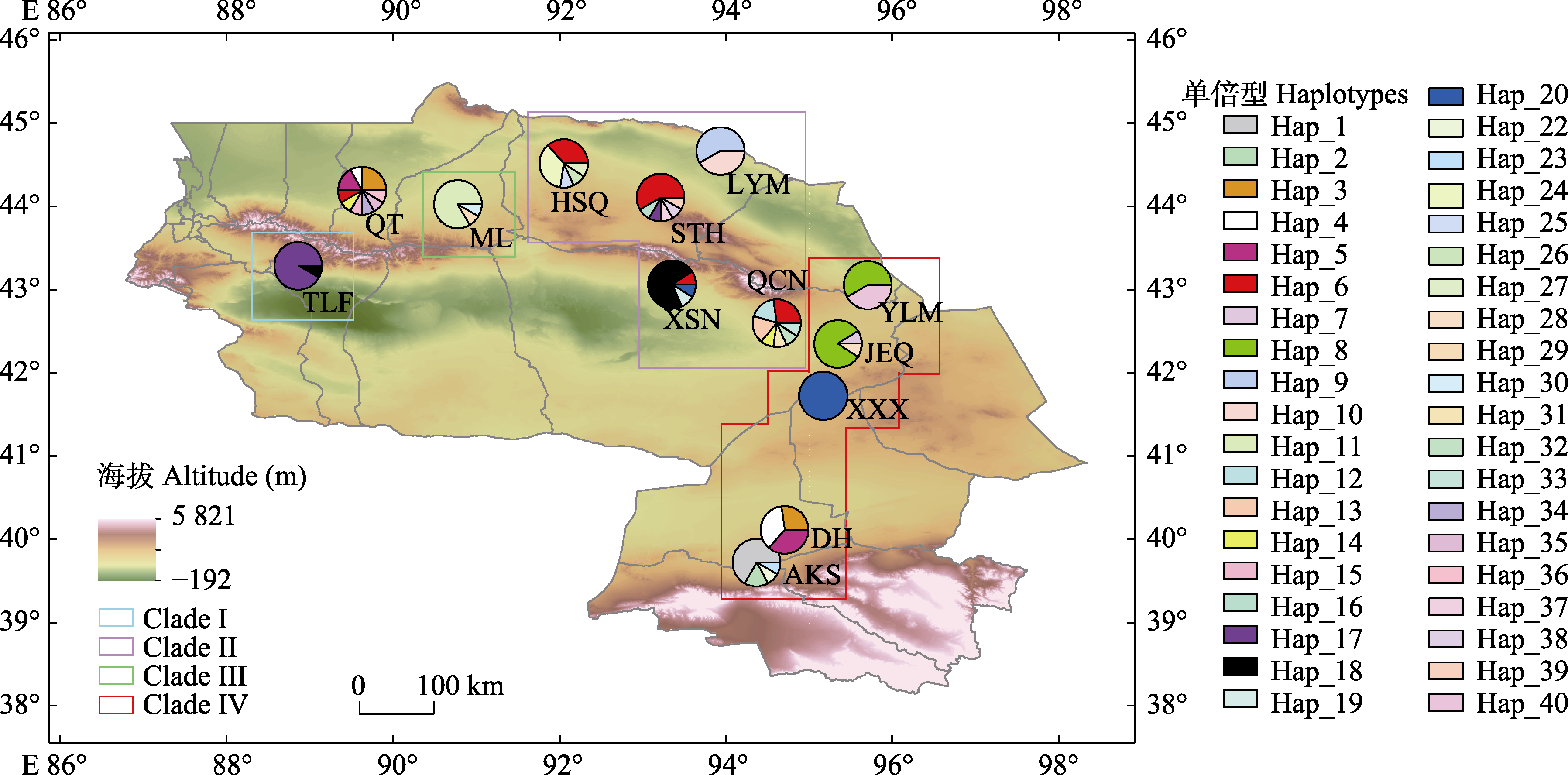
图4 梭梭13个种群的39个叶绿体单倍型的地理分布。图中缩写为13个采样种群编号, 详见表1。扇形图中不同颜色表示不同单倍型, 扇形大小表示该种群中某种单倍型的占比。Clade I-IV表示使用叶绿体基因组划分的4个类群。
Fig. 4 Geographical distribution of 39 haplotypes of 13 Haloxylon ammodendron populations. The abbreviations indicate population codes, as shown in Table 1. In the fan chart, the color represents different haplotypes, and the size of the fan represents the proportion of haplotypes in the population. Clade I-IV represent four clades divided according to chloroplast genomes.
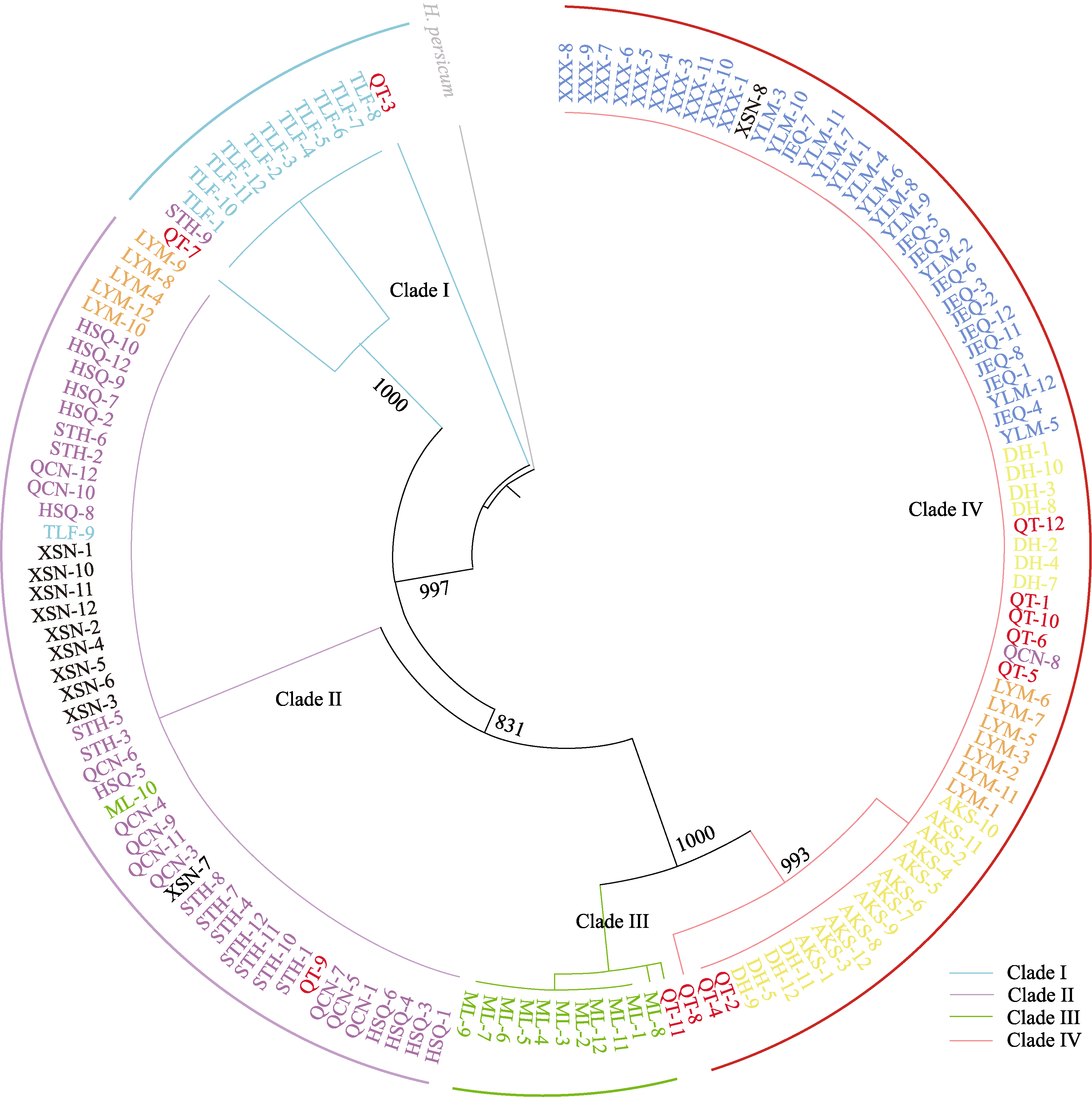
图5 基于cpDNA使用最大似然法构建149个梭梭个体最大似然系统发育树。每个枝条末端表示13个梭梭种群中149个个体, 奇台种群为红色, 吐鲁番种群为浅蓝色, 汉水泉、三塘湖、沁城乡种群为紫色, 老爷庙种群为橙色, 西山乡种群为黑色, 木垒种群为绿色, 镜儿泉、亚勒曼、星星峡种群为深蓝色, 阿克塞、敦煌种群为黄色, 各种群信息同表1。外围彩色条带(Clade I-IV)表示使用叶绿体基因将149个梭梭个体分为4个类群。
Fig. 5 A maximum likelihood phylogenetic tree of the 149 Haloxylon ammodendron based on chloroplast genomes. The end of each branch represents 149 individuals in 13 H. ammodendron populations, QT is red, TLF is light blue, HSQ, STH, QCN is purple, LYM is orange, XSN is brown, ML is green, JEQ, YLM, XXX is dark blue, AKS, DH is yellow. The information of populations is shown in Table 1. The outmost colored strips (Clade I-IV) indicate four clades divided according to chloroplast genomes.
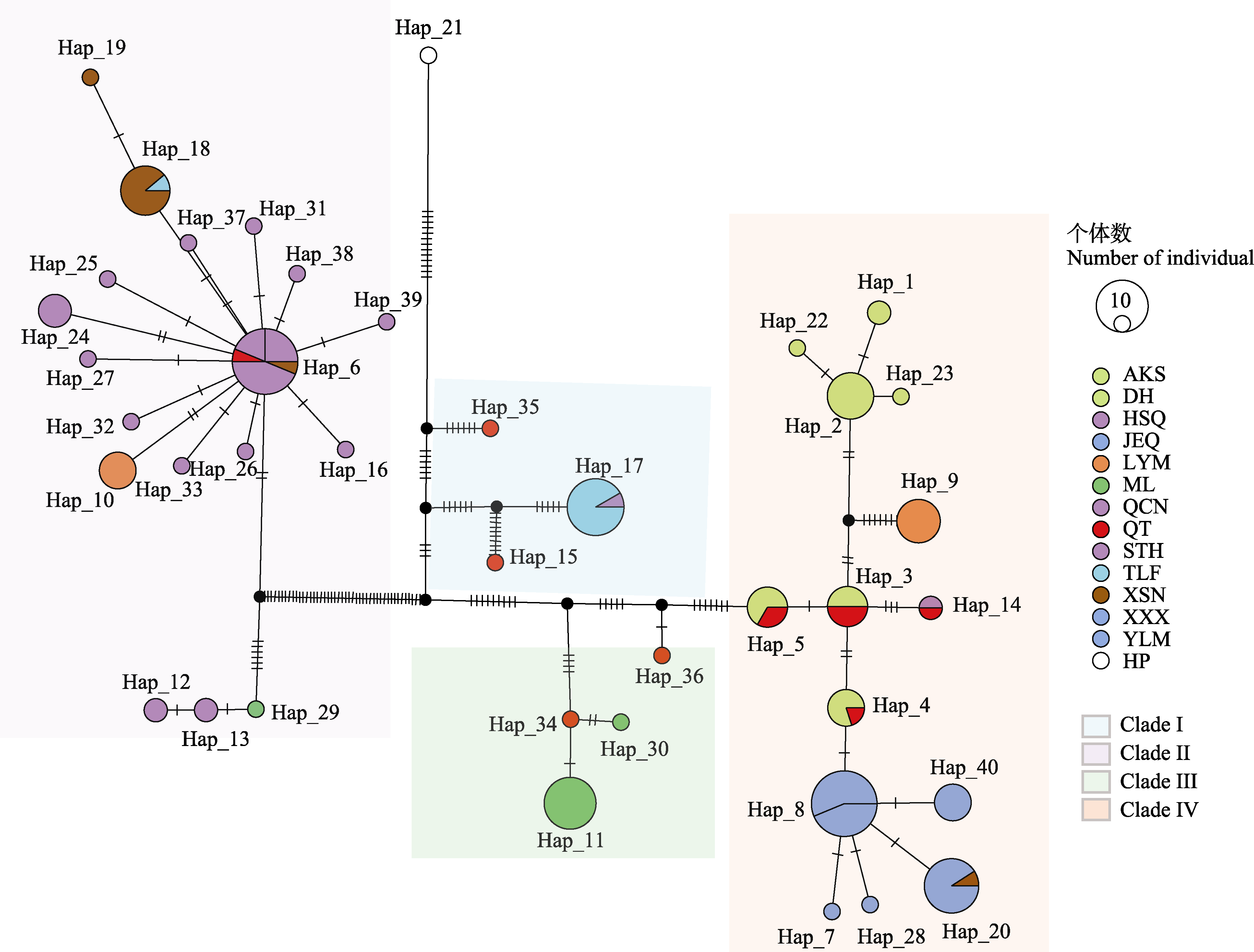
图6 基于cpDNA的单倍型(Hap)网络连接图。圆圈表示一种单倍型, 圆圈大小表示拥有该单倍型的个体数量, 不同颜色表示不同种群, 种群名称详见表1, 黑色圆圈表示未采样或已灭绝单倍型, 白色圆圈表示外类群白梭梭单倍型。扇形大小表示拥有某种单倍型的个体来自不同种群的占比。Clade I-IV表示使用叶绿体基因组划分的4个类群。
Fig. 6 Haplotype (Hap) network for the 39 haplotypes of Haloxylon ammodendron based on chloroplast genomes. The circles represent haplotypes, the size of the circle represents the number of individuals with the haplotype, and different colors represent different populations. The abbreviations indicate population codes, as shown in Table 1. The black circle represents the unsampled or extinct haplotypes, and the white circle represents the haplotype of Haloxylon persicum. The fan size indicates the proportion of individuals from different populations with a certain haplotype. Clade I-IV represent four clades divided according to chloroplast genomes.
| [1] |
Alexander DH, Novembre J, Lange K (2009). Fast model-based estimation of ancestry in unrelated individuals. Genome Research, 19, 1655-1664.
DOI PMID |
| [2] | Bai WN, Zhang DY (2014). Current status and future directions in plant phylogeography. Chinese Bulletin of Life Sciences, 26, 125-137. |
| [白伟宁, 张大勇 (2014). 植物亲缘地理学的研究现状与发展趋势. 生命科学, 26, 125-137.] | |
| [3] |
Chang CC, Chow CC, Tellier LC, Vattikuti S, Purcell SM, Lee JJ (2015). Second-generation PLINK: rising to the challenge of larger and richer datasets. GigaScience, 4, 7. DOI: 10.48550/arXiv.1410.4803.
PMID |
| [4] |
Chen H, Xie W, He H, Yu H, Chen W, Li J, Yu R, Yao Y, Zhang W, He Y, Tang X, Zhou F, Deng XW, Zhang Q (2014). A high-density SNP genotyping array for rice biology and molecular breeding. Molecular Plant, 7, 541-553.
DOI PMID |
| [5] | Chen Y, Ma S, Zhang D, Wei B, Huang G, Zhang Y, Ge B (2022). Diversification and historical demography of Haloxylon ammodendron in relation to Pleistocene climatic oscillations in northwestern China. PeerJ, 10, e14476. DOI: 10.7717/peerj.14476. |
| [6] |
Chen YT, Ma SM, Zhang D, Zhang L, Wang CC (2024). Diversity pattern and formation mechanism of sympatric Haloxylon ammodendron and Haloxylon persicum in Xinjiang, China. Chinese Journal of Plant Ecology, 48, 56-67.
DOI URL |
|
[陈雨婷, 马松梅, 张丹, 张林, 王春成 (2024). 新疆同域分布梭梭和白梭梭多样性格局及其形成机制. 植物生态学报, 48, 56-67.]
DOI |
|
| [7] | Cheng WM, Bao AM, Chai HX, Zhao SM, Fang Y (2018). Geomorphological Patterns and Effects in Xinjiang. Science Press, Beijing. |
| [程维明, 包安明, 柴慧霞, 赵尚民, 方月 (2018). 新疆地貌格局及其效应. 科学出版社, 北京.] | |
| [8] |
Danecek P, Auton A, Abecasis G, Albers CA, Banks E, DePristo MA, Handsaker RE, Lunter G, Marth GT, Sherry ST, McVean G, Durbin R (2011). The variant call format and VCFtools. Bioinformatics, 27, 2156-2158.
DOI PMID |
| [9] | Danecek P, Bonfield JK, Liddle J, Marshall J, Ohan V, Pollard MO, Whitwham A, Keane T, McCarthy SA, Davies RM, Li H (2021). Twelve years of SAMtools and BCFtools. GigaScience, 10, giab008. DOI: 10.1093/gigascience/giab008. |
| [10] | Dierckxsens N, Mardulyn P, Smits G (2017). NOVOPlasty: de novo assembly of organelle genomes from whole genome data. Nucleic Acids Research, 45, e18. DOI: 10.1093/nar/gkw955. |
| [11] | Dong ZB, Su ZZ, Qian GQ, Luo WY, Zhang ZS, Wu JF (2011). Geomorphology of the Kumtagh Desert. Science Press, Beijing. |
| [董治宝, 苏志珠, 钱广强, 罗万银, 张正偲, 吴晋峰 (2011). 库姆塔格沙漠风沙地貌. 科学出版社, 北京.] | |
| [12] |
Duan L, Fu L, Chen HF (2023). Phylogenomic cytonuclear discordance and evolutionary histories of plants and animals. Science China: Life Sciences, 66, 2946-2948.
DOI |
| [13] | Gavin DG, Fitzpatrick MC, Gugger PF, Heath KD, Rodríguez-Sánchez F, Dobrowski SZ, Hampe A, Hu FS, Ashcroft MB, Bartlein PJ, Blois JL, Carstens BC, Davis EB, de Lafontaine G, Edwards ME, et al. (2014). Climate refugia: joint inference from fossil records, species distribution models and phylogeography. New Phytologist, 204, 37-54. |
| [14] | Guo F, Wang Q (2013). A summary of the desert formation and evolution in West China I. Journal of Foshan University (Natural Science Edition), 31(1), 56-61. |
| [郭峰, 王亲 (2013). 中国西部干旱区沙漠形成演化综述I. 佛山科学技术学院学报(自然科学版), 31(1), 56-61.] | |
| [15] | Guo QS, Wang CL, Guo ZH, Tan DY, Shi ZM (2005). Geographic distribution of existing Haloxylon desert vegetation and its patch character in China. Scientia Silvae Sinicae, 41(5), 2-7. |
| [郭泉水, 王春玲, 郭志华, 谭德远, 史作民 (2005). 我国现存梭梭荒漠植被地理分布及其斑块特征. 林业科学, 41(5), 2-7.] | |
| [16] | Hou XY (2001). Vegetation Atlas of China. Science Press, Beijing. |
| [侯学煜 (2001). 中国植被图集. 科学出版社, 北京.] | |
| [17] |
Hu Y, Wang X, Zhang XX, Zhou W, Chen XY, Hu XS (2019). Advancing phylogeography with chloroplast DNA markers. Biodiversity Science, 27, 219-234.
DOI |
|
[胡颖, 王茜, 张新新, 周玮, 陈晓阳, 胡新生 (2019). 叶绿体DNA标记在谱系地理学中的应用研究进展. 生物多样性, 27, 219-234.]
DOI |
|
| [18] | Jia SW, Zhang ML (2021). Introgression of phylogeography lineages of Convolvulus gortschakovii (Convolvulaceae) in the northwest China. Plant Systematics and Evolution, 307, 19. DOI: 10.1007/s00606-020-01734-z. |
| [19] | Krutovsky KV, Neale DB (2005). Nucleotide diversity and linkage disequilibrium in cold-hardiness- and wood- quality-related candidate genes in Douglas fir. Genetics, 171, 2029-2041. |
| [20] |
Li H, Durbin R (2009). Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics, 25, 1754-1760.
DOI PMID |
| [21] |
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R (2009). The sequence alignment/map format and SAMtools. Bioinformatics, 25, 2078-2079.
DOI PMID |
| [22] | Li Q (2020). Solar forcing of desert vegetation and drought frequency during the last 2700 years in the interior Qaidam Basin, northeastern Tibetan Plateau. Scientia Sinica (Terrae), 50, 995-1008. |
| [李泉 (2020). 过去2700年太阳活动对青藏高原东北部柴达木盆地荒漠植被与气候干旱频率的驱动. 中国科学: 地球科学, 50, 995-1008.] | |
| [23] | Li SJ, Wang ZH, Su PX, Wang H, Wang FX, Cui LJ (2022). Research progress on the trade-off strategy and functional diversity of desert plants. Acta Ecologica Sinica, 42, 7308-7320. |
| [李善家, 王子濠, 苏培玺, 王辉, 王福祥, 崔莉娟 (2022). 荒漠植物性状权衡策略及功能多样性研究进展. 生态学报, 42, 7308-7320.] | |
| [24] |
Liu YK, Zhu ZW, Chen L, Zou J, Tong HW, Zhu G, He WJ, Zhang YQ, Gao CB (2020). Revealing the genetic diversity of wheat varieties (lines) in China based on SNP markers. Acta Agronomica Sinica, 46, 307-314.
DOI URL |
|
[刘易科, 朱展望, 陈泠, 邹娟, 佟汉文, 朱光, 何伟杰, 张宇庆, 高春保 (2020). 基于SNP标记揭示我国小麦品种(系)的遗传多样性. 作物学报, 46, 307-314.]
DOI |
|
| [25] |
Ma SM, Zhang ML (2012). Phylogeography and conservation genetics of the relic Gymnocarpos przewalskii (Caryophyllaceae) restricted to northwestern China. Conservation Genetics, 13, 1531-1541.
DOI URL |
| [26] | Sang YP, Long ZQ, Dan XM, Feng JJ, Shi TT, Jia CF, Zhang XX, Lai Q, Yang GL, Zhang HY, Xu XT, Liu HH, Jiang YZ, Ingvarsson PK, Liu JQ, et al. (2022). Genomic insights into local adaptation and future climate-induced vulnerability of a keystone forest tree in East Asia. Nature Communications, 13, 6541. DOI: 10.1038/s41467-022-34206-8. |
| [27] |
Simonsen KL, Churchill GA, Aquadro CF (1995). Properties of statistical tests of neutrality for DNA polymorphism data. Genetics, 141, 413-429.
DOI PMID |
| [28] | Sturm AB, Eckert RJ, Méndez JG, González-Díaz P, Voss JD (2020). Population genetic structure of the great star coral, Montastraea cavernosa, across the Cuban archipelago with comparisons between microsatellite and SNP markers. Scientific Reports, 10, 15432. DOI: 10.1038/s41598-020-72112-5. |
| [29] | Sun FF, Nie YB, Ma SM, Wei B, Ji WQ (2019). Species differentiation of Haloxylon ammodendron and Haloxylon persicum based on its and cpDNA sequences. Scientia Silvae Sinicae, 55(3), 43-53. |
| [孙芳芳, 聂迎彬, 马松梅, 魏博, 吉万全 (2019). 基于ITS和cpDNA序列的梭梭和白梭梭物种分化. 林业科学, 55(3), 43-53.] | |
| [30] |
Suo ZL, Jia ZQ, Lu Q, Pan BR, Jin XB, Xu G, Peng XQ, Sun HB, Tao YH (2012). Distinguishing Haloxylon persicum and H. ammodendron (Haloxylon Bunge, Amaranthaceae) using DNA marker. AASRI Procedia, 1, 305-310.
DOI URL |
| [31] | Wang M, Zhang L, Tong S, Jiang D, Fu Z (2022). Chromosome-level genome assembly of a xerophytic plant, Haloxylon ammodendron. DNA Research, 29, dsac006. DOI: 10.1093/dnares/dsac006. |
| [32] | Wei B (2020). Genetic Characteristics and Conservation Strategies of Haloxylon ammodendron Populations in Desert. Master degree dissertation, Shihezi University, Shihezi, Xinjiang. |
| [魏博 (2020). 荒漠植物梭梭居群的遗传特征及保护策略研究. 硕士学位论文, 石河子大学, 新疆石河子.] | |
| [33] | Wu ZY (1980). Vegetation of China. Science Press, Beijing. |
| [吴征镒 (1980). 中国植被. 科学出版社, 北京.] | |
| [34] | Yin HX, Wang LR, Shi Y, Qian CJ, Zhou HK, Wang WY, Ma XF, Tran LP, Zhang BY (2020). The east Asian winter monsoon acts as a major selective factor in the intraspecific differentiation of drought-tolerant Nitraria tangutorum in northwest China. Plants, 9, 1100. DOI: 10.3390/plants9091100. |
| [35] | Zhang D, Ma SM, Wei B, Wang CC, Zhang L, Yan H (2022). Historical distribution pattern and driving mechanism of Haloxylon in China. Biodiversity Science, 30, 42-51. |
| [张丹, 马松梅, 魏博, 王春成, 张林, 闫涵 (2022). 中国梭梭属植物历史分布格局及其驱动机制. 生物多样性, 30, 42-51.] | |
| [36] | Zhang P, Dong YZ, Wei Y, Hu CZ (2006). ISSR analysis of genetic diversity of Haloxylon ammodendron (C. A. Mey.) bunge in Xinjiang. Acta Botanica Boreali-Occidentalia Sinica, 26, 1337-1341. |
| [张萍, 董玉芝, 魏岩, 胡成志 (2006). 利用ISSR标记对新疆梭梭遗传多样性的研究. 西北植物学报, 26, 1337-1341.] | |
| [37] | Zhu SY, Wu HB, Li Q, Zhang CX, Yu YY, Sun AZ, Guo ZT (2016). Aridification in northwestern China since the late Cenozoic evidenced by the vegetation change. Quaternary Sciences, 36, 820-831. |
| [祝淑雅, 吴海斌, 李琴, 张春霞, 于严严, 孙爱芝, 郭正堂 (2016). 晚新生代以来中国西北植被演化及反映的干旱化过程. 第四纪研究, 36, 820-831.] |
| [1] | 韩雨晴, 熊伟, 吴波, 卢琦, 杨文斌, 刘雅莉, 张景波, 辛智鸣, 马迎宾, 廉泓林, 王思涵. 乌兰布和沙漠梭梭茎干液流对降雨脉冲的响应[J]. 植物生态学报, 2024, 48(9): 1172-1179. |
| [2] | 马佳正, 陈雨婷, 马松梅, 张丹, 贺凌云. 基于多源数据的新疆沙漠植物白梭梭遗传格局与扩散路径模拟[J]. 植物生态学报, 2024, 48(10): 1326-1335. |
| [3] | 陈雨婷, 马松梅, 张丹, 张林, 王春成. 新疆同域分布梭梭和白梭梭多样性格局及其形成机制[J]. 植物生态学报, 2024, 48(1): 56-67. |
| [4] | 陈图强, 徐贵青, 刘深思, 李彦. 干旱胁迫下梭梭水力性状调整与非结构性碳水化合物动态[J]. 植物生态学报, 2023, 47(10): 1407-1421. |
| [5] | 张利锐, 彭艳玲, 任广朋, 周永锋, 李忠虎, 刘建全. 马尾松和黄山松两个核基因位点的群体遗传多样性和种间分化[J]. 植物生态学报, 2011, 35(5): 531-538. |
| [6] | 严巧娣, 苏培玺, 陈宏彬, 张岭梅. 五种C4荒漠植物光合器官中含晶细胞的比较分析[J]. 植物生态学报, 2008, 32(4): 873-882. |
| [7] | 苏培玺, 安黎哲, 马瑞君, 刘新民. 荒漠植物梭梭和沙拐枣的花环结构及C4光合特征[J]. 植物生态学报, 2005, 29(1): 1-7. |
| [8] | 侯天侦, 梁远强. 新疆甘家湖梭梭林的光合、水分生理生态的研究[J]. 植物生态学报, 1991, 15(2): 141-150. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||
Copyright © 2026 版权所有 《植物生态学报》编辑部
地址: 北京香山南辛村20号, 邮编: 100093
Tel.: 010-62836134, 62836138; Fax: 010-82599431; E-mail: apes@ibcas.ac.cn, cjpe@ibcas.ac.cn
备案号: 京ICP备16067583号-19
![]()