
Chin J Plant Ecol ›› 2013, Vol. 37 ›› Issue (8): 750-757.DOI: 10.3724/SP.J.1258.2013.00078
Special Issue: 生物多样性
• Research Articles • Previous Articles Next Articles
Received:2013-02-05
Accepted:2013-05-15
Online:2013-02-05
Published:2013-08-07
Contact:
WANG Ai-Li
WANG Ai-Li. Microbial community diversity in the rhizosphere of wetland plants examined by phospholipid fatty acid and polymerase chain reaction denaturing gradient gel electrophoresis[J]. Chin J Plant Ecol, 2013, 37(8): 750-757.
Add to citation manager EndNote|Ris|BibTeX
URL: https://www.plant-ecology.com/EN/10.3724/SP.J.1258.2013.00078
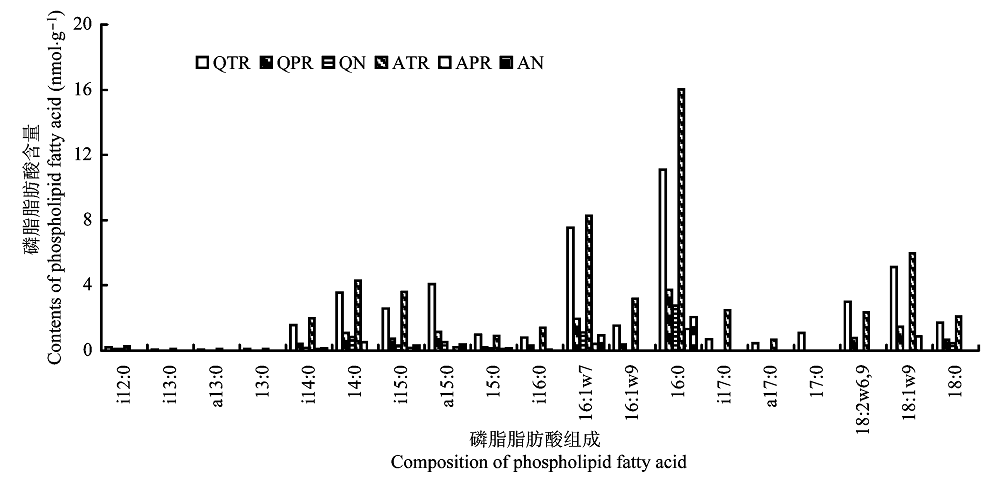
Fig. 1 Composition of phospholipid fatty acid in sediment samples. A, Aiwan Lake; N, non-rhizosphere; PR, Phragmites australis rhizosphere; Q, Qingnian Lake; TR, Typha orientalis rhizosphere.
| 青年湖 Qingnian Lake | 爱晚湖 Aiwan Lake | ||||||
|---|---|---|---|---|---|---|---|
| 东方香蒲根际 Typha orientalis rhizosphere | 芦苇根际 Phragmites australis rhizosphere | 非根际 Non- rhizosphere | 东方香蒲根际 Typha orientalis rhizosphere | 芦苇根际 Phragmites australis rhizosphere | 非根际 Non- rhizosphere | ||
| 微生物生物量 Microbial biomass | 44.17B (2.19) | 12.35C (1.48) | 5.99D (0.79) | 51.17A (3.19) | 3.52E (0.25) | 3.93E (0.27) | |
| 革兰氏阳性细菌 Gram-positive bacteria | 8.65A (0.85) | 2.17B (0.12) | 2.10C (0.08) | 26.40A (2.30) | 1.86C (0.06) | 1.78C (0.05) | |
| 革兰氏阴性细菌 Gram-negative bacteria | 14.21B (0.13) | 3.81C (0.05) | - | 2.35A (0.22) | - | - | |
| 阳性菌与阴性菌的比值 Ratio of gram-positive bacteria to gram-negative bacteria | 0.61B (0.05) | 0.57B (0.03) | 0.83C (0.02) | 8.11A (0.75) | 0.46E (0.01) | 0.70D (0.01) | |
| 细菌生物量 Bacterial biomass | 24.91A (2.15) | 6.20B (0.52) | 1.12E (0.03) | 17.39A (1.45) | 1.30D (0.04) | 0.93F (0.02) | |
| 真菌生物量 Fungal biomass | 2.99A (0.25) | 0.77B (0.05) | 0.74A (0.0.2) | 0.47C (0.01) | 0.35D (0.01) | 0.75A (0.03) | |
Table 1 Contents of phospholipid fatty acids in sediments (nmol·g-1 dry weight)
| 青年湖 Qingnian Lake | 爱晚湖 Aiwan Lake | ||||||
|---|---|---|---|---|---|---|---|
| 东方香蒲根际 Typha orientalis rhizosphere | 芦苇根际 Phragmites australis rhizosphere | 非根际 Non- rhizosphere | 东方香蒲根际 Typha orientalis rhizosphere | 芦苇根际 Phragmites australis rhizosphere | 非根际 Non- rhizosphere | ||
| 微生物生物量 Microbial biomass | 44.17B (2.19) | 12.35C (1.48) | 5.99D (0.79) | 51.17A (3.19) | 3.52E (0.25) | 3.93E (0.27) | |
| 革兰氏阳性细菌 Gram-positive bacteria | 8.65A (0.85) | 2.17B (0.12) | 2.10C (0.08) | 26.40A (2.30) | 1.86C (0.06) | 1.78C (0.05) | |
| 革兰氏阴性细菌 Gram-negative bacteria | 14.21B (0.13) | 3.81C (0.05) | - | 2.35A (0.22) | - | - | |
| 阳性菌与阴性菌的比值 Ratio of gram-positive bacteria to gram-negative bacteria | 0.61B (0.05) | 0.57B (0.03) | 0.83C (0.02) | 8.11A (0.75) | 0.46E (0.01) | 0.70D (0.01) | |
| 细菌生物量 Bacterial biomass | 24.91A (2.15) | 6.20B (0.52) | 1.12E (0.03) | 17.39A (1.45) | 1.30D (0.04) | 0.93F (0.02) | |
| 真菌生物量 Fungal biomass | 2.99A (0.25) | 0.77B (0.05) | 0.74A (0.0.2) | 0.47C (0.01) | 0.35D (0.01) | 0.75A (0.03) | |
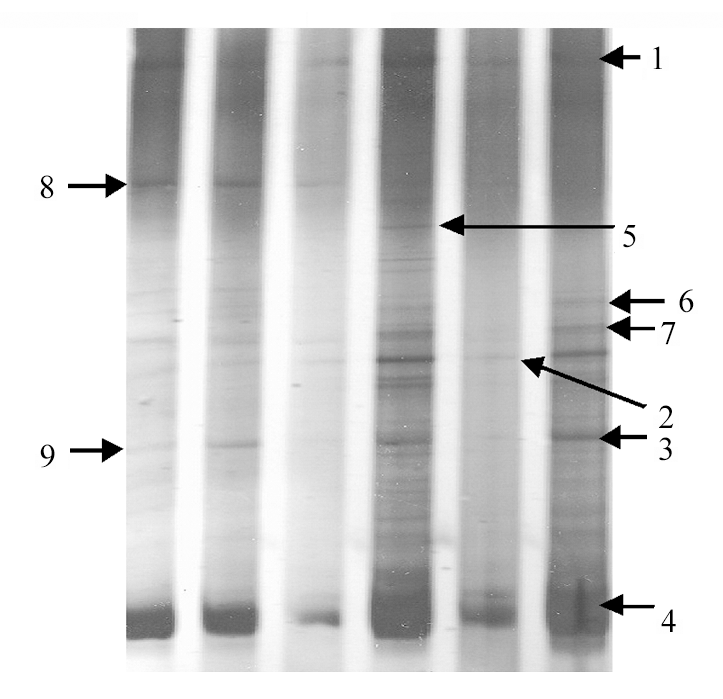
Fig. 2 Denaturing gradient gel electrophoresis separation of different sediment samples. From left to right are Typha orientalis rhizosphere in Qingnian Lake, Phragmites australis rhizosphere in Qingnian Lake, non-rhizosphere in Qingnian Lake, Typha orientalis rhizosphere in Aiwan Lake, non-rhizosphere in Aiwan Lake, Phragmites australis rhizosphere in Aiwan Lake, respectively. The numbers in figure refer to the number of bands.
| 青年湖 Qingnian Lake | 爱晚湖 Aiwan Lake | ||||||
|---|---|---|---|---|---|---|---|
| 东方香蒲根际 Typha orientalis rhizosphere | 芦苇根际 Phragmites australis rhizosphere | 非根际 Non- rhizosphere | 东方香蒲根际 Typha orientalis rhizosphere | 芦苇根际 Phragmites australis rhizosphere | 非根际 Non- rhizosphere | ||
| 磷脂脂肪酸 Phospholipid fatty acids | 2.39A (0.20) | 2.20A (0.18) | 1.72B (0.15) | 2.25A (0.18) | 1.79B (0.12) | 1.35C (0.09) | |
| 变性梯度凝胶电泳 Denaturing gradient gel electrophoresis | 2.46A (0.21) | 2.32A (0.19) | 1.68B (0.15) | 2.87A (0.22) | 2.61A (0.20) | 1.66B (0.13) | |
Table 2 Diversity index of microbial community in different sediments
| 青年湖 Qingnian Lake | 爱晚湖 Aiwan Lake | ||||||
|---|---|---|---|---|---|---|---|
| 东方香蒲根际 Typha orientalis rhizosphere | 芦苇根际 Phragmites australis rhizosphere | 非根际 Non- rhizosphere | 东方香蒲根际 Typha orientalis rhizosphere | 芦苇根际 Phragmites australis rhizosphere | 非根际 Non- rhizosphere | ||
| 磷脂脂肪酸 Phospholipid fatty acids | 2.39A (0.20) | 2.20A (0.18) | 1.72B (0.15) | 2.25A (0.18) | 1.79B (0.12) | 1.35C (0.09) | |
| 变性梯度凝胶电泳 Denaturing gradient gel electrophoresis | 2.46A (0.21) | 2.32A (0.19) | 1.68B (0.15) | 2.87A (0.22) | 2.61A (0.20) | 1.66B (0.13) | |
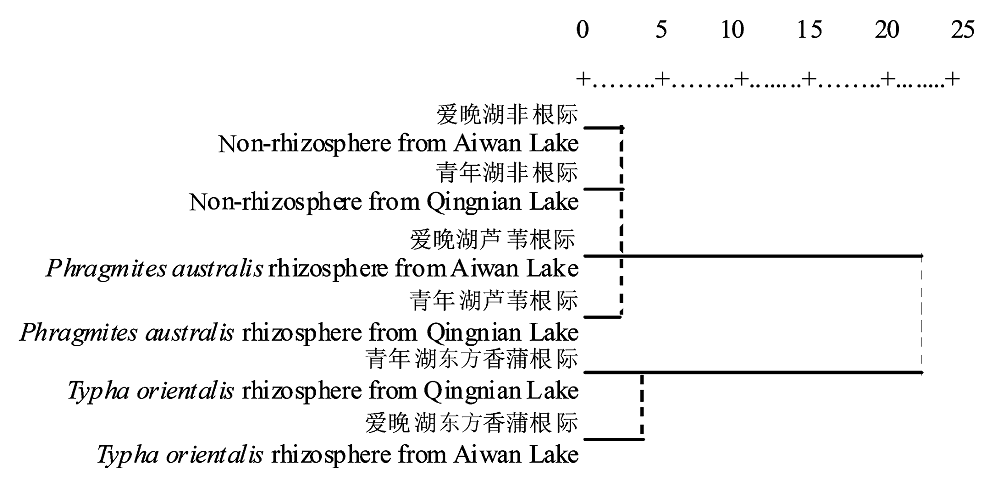
Fig. 3 Cluster analysis of phospholipid fatty acid data of microbial community from sediment samples. The numbers in figure refer to average distance between the groups in cluster analysis and the unit of the number is centimeter.
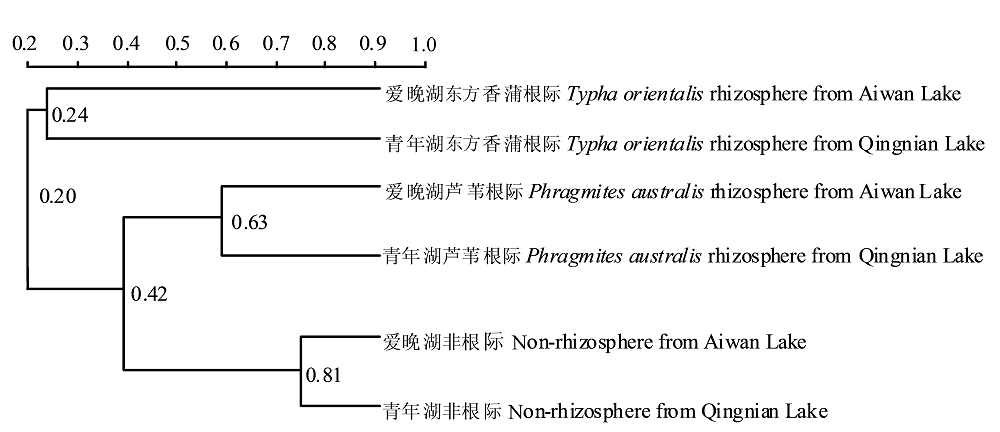

Fig. 4 Cluster analysis of denaturing gradient gel electrophoresis bands of microbial community from sediment samples. The numbers in figure refer to the similarity among denaturing gradient gel electrophoresis bands between the groups in cluster analysis. For example, 0.81 refers to 81% of similarity.
| [1] |
Berg G, Roskot N, Steidle A, Eberl L, Zock A, Smalla K (2002). Plant-dependent genotypic and phenotypic diver- sity of antagonistic rhizobacteria isolated from different Verticillium host plants. Applied and Environmental Microbiology, 68, 3328-3338.
URL PMID |
| [2] | Chi J, Yang R, Wang AL (2012). Effect of wetland plant species and growth strategy on the distribution of PAEs and their monoester metabolites in the rhizosphere. Journal of Lake Science, 24, 416-421. (in Chinese with English abstract) |
| [ 迟杰, 杨瑞, 王爱丽 (2012). 湿地植物种类和生长方式对根际酞酸酯及其单酯代谢物分布特征的影响. 湖泊科学, 24, 416-421.] | |
| [3] |
Dong XL, Reddy GB (2010). Soil bacterial communities in constructed wetlands treated with swine wastewater using PCR-DGGE technique. Bioresource Technology, 101, 1175-1182.
DOI URL PMID |
| [4] |
Fan SX, Li PJ, Gong ZQ, Ren WX, He N (2008). Promotion of pyrene degradation in rhizosphere of alfalfa (Medicago sativa L.). Chemosphere, 71, 1593-1598.
DOI URL PMID |
| [5] | Haberl R, Grego S, Langergraber G, Kadlec RH, Cicalini AR, Dias SM, Novais JM, Aubert S, Gerth A, Thomas H, Hebner A (2003). Constructed wetlands for the treatment of organic pollutants. Journal of Soils and Sediments, 3, 109-124. |
| [6] |
Hallberg KB, Johnson BD (2005). Microbiology of a wetland ecosystem constructed to remediate mine drainage from a heavy metal mine. Science of the Total Environment, 338, 53-66.
DOI URL PMID |
| [7] | Heinrich D, Hess D (1985). Chemotactic attraction of Azospirillum lipoferum by wheat roots and characterization of some attractants. Canadian Journal of Microbiology, 31, 26-31. |
| [8] |
Iwamoto T, Tani K, Nakamura K, Suzuki Y, Kitagawa M, Equchi M, Nasu M (2000). Monitoring impact of in suit biostimulation treatment on ground water bacterial community by DGGE. FEMS Microbiology Ecology, 32, 129-141.
DOI URL PMID |
| [9] |
Jenkins MB, Lion LW (1993). Mobile bacteria and transport of poly-nuclear aromatic hydrocarbons in porous media. Applied and Environmental Microbiology, 59, 3306-3313.
DOI URL PMID |
| [10] |
Li M, Zhou QH, Tao M, Wang Y, Jiang LJ, Wu ZB (2010). Comparative study of microbial community structure in different filter media of constructed wetland. Journal of Environmental Sciences, 22, 127-133.
DOI URL |
| [11] | Li RH, Guan YT, He M, Hu HY, Jiang ZP (2006). Pilot- scale study on riparian mixed plant zones treating polluted river water. Environmental Science, 27, 651-654 (in Chinese with English abstract) |
| [ 李睿华, 管运涛, 何苗, 胡洪营, 蒋展鹏 (2006). 河岸混合植物带处理受污染河水中试研究. 环境科学, 27, 651-654.] | |
| [12] |
Liu M, Yang Y, Xu S, Liu H, Hou L, Ou D, Liu Q, Cheng S (2006). HCHs and DDTs in salt marsh plants (Scirpus) from the Yangtze estuary and nearby coastal areas, China. Chemosphere, 62, 440-448.
DOI URL PMID |
| [13] |
Miglioranza KSB, de Moreno JEA, Moreno VJ (2004). Organochlorine pesticides sequestered in the aquatic macrophyte Schoenoplectus californicus (C.A. Meyer) Soják from a shallow lake in Argentina. Water Research, 38, 1765-1772.
DOI URL PMID |
| [14] |
Ruiz-Rueda O, Hallin S, Baneras L (2009). Structure and function of denitrifying and nitrifying bacteria communities in relation to the plant species in a constructed wetland. FEMS Microbiology Ecology, 67, 308-319.
DOI URL PMID |
| [15] |
Sirivedhim T, Gray KA (2006). Factors affecting denitrification rates in experimental wetlands: field and laboratory studies. Ecological Engineering, 26, 167-181.
DOI URL |
| [16] |
Smalla K, Wieland G, Buchner A, Zock A, Parzy J, Kaiser S, Roskot N, Heuer H, Berg G (2001). Bulk and rhizosphere soil bacterial communities studied by denaturing gradient gel electrophoresis: plant-dependent enrichment and seasonal shifts revealed. Applied and Environmental Microbiology, 67, 4742-4751.
URL PMID |
| [17] |
Tawney L, Becker JG, Baldwin AH (2008). A novel dual-compartment, continuous-flow wetland microcosm to assess cis-dichloroethene removal from the rhizosphere. International Journal of Phytoremediation, 10, 455-471.
URL PMID |
| [18] |
Toyama T, Sato Y, Inoue D, Sei K, Chang YC, Kikuchi S, Ike M (2009) Biodegradation of bisphenol A and bisphenol F in the rhizosphere sediment of Phragmites australis. Journal of Bioscience and Bioengineering, 108, 147-150.
DOI URL PMID |
| [19] | Wang AL (2011). Dissipation Behaviors of Phthalic Acid Esters in the Rhizosphere of Wetland Plants. PhD dissertation, Tianjin University, Tianjin. (in Chinese with English abstract) |
| [ 王爱丽 (2011). 酞酸酯在湿地植物根际环境中的消减行为. 博士学位论文, 天津大学, 天津.] | |
| [20] | Xiang XM, Song CX, Li YS, Sun XY (2004). Microorganism features of Typha Latifolia and Phragmites Australis at rhizosphere. Chinese Journal of Environmental Protection Science, 30, 35-38. (in Chinese with English abstract) |
| [ 项学敏, 宋春霞, 李彦生, 孙祥宇 (2004). 湿地植物芦苇和东方香蒲根际微生物特性研究. 环境保护科学, 30, 35-38.] | |
| [21] | Xu XL, Lu XX, Lei XD, Cao LK (2012). Effects of hydrophytes on removal of nitrogen and phosphorus in eutrophic water. Journal of Shanghai Jiaotong University (Agricultural Science), 30, 8-14. (in Chinese with English abstract) |
| [ 徐秀玲, 陆欣欣, 雷先德, 曹林奎 (2012). 不同水生植物对富营养化水体中氮磷去除效果的比较. 上海交通大学学报(农业科学版), 30, 8-14.] | |
| [22] |
Xue D, Yao HY, Ge DY, Huang CY (2008). Soil microbial community structure in diverse land use systems: a comparative study using Biolog, DGGE, and PLFA analyses. Pedosphere, 18, 653-663.
DOI URL |
| [23] | Yang R (2011) The Studies on Microbial Ecology of the Rhizosphere of Emergent Plants. Master degree dissertation, Tianjin University, Tianjin. (in Chinese with English abstract) |
| [ 杨瑞. 挺水植物根际微生物生态研究. 硕士学位论文, 天津大学, 天津.] | |
| [24] |
Zelles L (1997). Phospholipid fatty acid profiles in selected members of soil microbial communities. Chemosphere, 35, 275-294.
DOI URL PMID |
| [25] |
Zhang QR, Zhou QX, Ren LP, Zhu YG, Sun SL (2006). Ecological effects of crude oil residues on the functional diversity of soil microorganisms in three weed rhizospheres. Journal of Environmental Sciences, 18, 1101-1106.
DOI URL |
| [26] | Zhao XG (2011). Effect of Emergent Plants on the Circulation of Nitrogen and Phosphorus of Eutrophic Lakes. Master degree dissertation, Tianjin University, Tianjin. (in Chinese with English abstract) |
| [ 赵旭光. 挺水植物对富营养化湖泊水体中氮磷循环的影响. 硕士学位论文, 天津大学, 天津.] |
| [1] | HU Zhao-Yi, CHEN Tian-Song, ZHAO Li, XU Pei-Xuan, WU Zheng-Jiang, DONG Li-Qin, ZHANG Kun. Effect of water level drop on nitrogen and phosphorus reabsorption of Carex muliensis in a herb swamp in Zoigê wetland, China [J]. Chin J Plant Ecol, 2023, 47(6): 847-855. |
| [2] | ZHANG Qi, FENG Ke, CHANG Zhi-Hui, HE Shuang-Hui, XU Wei-Qi. Effects of shrub encroachment on plant and soil microbial in the forest-grassland ecotone [J]. Chin J Plant Ecol, 2023, 47(6): 770-781. |
| [3] | FENG Ke, LIU Dong-Mei, ZHANG Qi, AN Jing, HE Shuang-Hui. Effect of tourism disturbance on soil microbial diversity and community structure in a Pinus tabuliformis forest [J]. Chin J Plant Ecol, 2023, 47(4): 584-596. |
| [4] | XU Gan-Jun, WU Sheng-Yi, LI Wei, ZHAO Xin-Sheng, NIE Lei-Chao, TANG Xi-Ying, ZHAI Xia-Jie. Estimation of carbon storage in Shaanxi Yellow River Wetland Provincial Nature Reserve [J]. Chin J Plant Ecol, 2023, 47(4): 469-478. |
| [5] | WANG Wen-Wei, HAN Wei-Peng, LIU Wen-Wen. Short-term response of leaf functional traits of the invasive plant Spartina alterniflora to a tidal gradient in coastal wetlands [J]. Chin J Plant Ecol, 2023, 47(2): 216-226. |
| [6] | WANG Jun-Qiang, LIU Bin, CHANG Feng, MA Zi-Jing, FAN Jia-Hui, HE Xiang-Ju, YOU Si-Xue, Aerziguli ABUDUREXITI, YANG Ying-Ke, SHEN Xin-Yan. Plant functional traits and ecological stoichiometric characteristics under water-salt gradient in the lakeshore zone of Bosten Lake [J]. Chin J Plant Ecol, 2022, 46(8): 961-970. |
| [7] | ZENG Kai-Na, SUN Hao-Ran, SHEN Yi-Chun, REN Ming-Xun. Pollination network and seasonal dynamics of Yangshan Wetland in Hainan Island, China [J]. Chin J Plant Ecol, 2022, 46(7): 775-784. |
| [8] | WEN Ke, YAO Huan-Mei, GONG Zhu-Qing, NA Ze-Lin, WEI Yi-Ming, HUANG Yi, CHEN Hua-Quan, LIAO Peng-Ren, TANG Li-Ping. Influence of inundation frequency change on enhanced vegetation index of wetland vegetation in Poyang Lake, China [J]. Chin J Plant Ecol, 2022, 46(2): 148-161. |
| [9] | NIE Xiu-Qing, WANG Dong, ZHOU Guo-Ying, XIONG Feng, DU Yan-Gong. Soil microbial biomass carbon, nitrogen, phosphorus and their stoichiometric characteristics in alpine wetlands in the Three Rivers Sources Region [J]. Chin J Plant Ecol, 2021, 45(9): 996-1005. |
| [10] | JIANG Xin, NIU Ke-Chang. Effects of grass mixed-sowing on soil microbial diversity on the Qingzang (Tibetan) Plateau [J]. Chin J Plant Ecol, 2021, 45(5): 539-551. |
| [11] | XUE Peng-Fei, LI Wen-Long, ZHU Gao-Feng, ZHOU Hua-Kun, LIU Chen-Li, YAN He-Piao. Changes in the pattern of an alpine wetland landscape in Maqu County in the first meander of the Yellow River [J]. Chin J Plant Ecol, 2021, 45(5): 467-475. |
| [12] | PEI Guang-Ting, SUN Jian-Fei, HE Tong-Xin, HU Bao-Qing. Effects of long-term human disturbances on soil microbial diversity and community structure in a karst grassland ecosystem of northwestern Guangxi, China [J]. Chin J Plant Ecol, 2021, 45(1): 74-84. |
| [13] | YU Liang, LI Jun-Li, BAO An-Ming, BAI Jie, HUANG Yue, LIU Tie, SHEN Zhan-Feng. Temporal areal changes of wetlands in the lower reaches of the Tarim River and their responses to ecological water conveyance [J]. Chin J Plant Ecol, 2020, 44(6): 616-627. |
| [14] | CHEN Yu-Han, LUO Yi-Fu, SUN Xin-Sheng, WEI Guan-Wen, HUANG Wen-Jun, LUO Fang-Li, YU Fei-Hai. Effects of waterlogging and increased soil nutrients on growth and reproduction of Polygonum hydropiper in the hydro-fluctuation belt of the Three Gorges Reservoir Region [J]. Chin J Plant Ecol, 2020, 44(11): 1184-1194. |
| [15] | YAN Peng-Fei, ZHAN Peng-Fei, XIAO De-Rong, WANG Yi, YU Rui, LIU Zhen-Ya, WANG Hang. Effects of simulated warming and decomposition interface on the litter decomposition rate of Zizania latifolia and its phyllospheric microbial community structure and function [J]. Chin J Plant Ecol, 2019, 43(2): 107-118. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2026 Chinese Journal of Plant Ecology
Tel: 010-62836134, 62836138, E-mail: apes@ibcas.ac.cn, cjpe@ibcas.ac.cn
![]()
